Welcome to
RBP World!
Functions and Diseases
| RBP Type | Canonical_RBPs |
| Diseases | CancerSARS-COV-2DengueZika |
| Drug | N.A. |
| Main interacting RNAs | mRNA |
| Moonlighting functions | N.A. |
| Localizations | Stress granucle |
| BulkPerturb-seq | DataSet_01_143 DataSet_01_46 |
Description
| Ensembl ID | ENSG00000153944 | Gene ID | 124540 | Accession | 18585 |
| Symbol | MSI2 | Alias | MSI2H | Full Name | musashi RNA binding protein 2 |
| Status | Confidence | Length | 428839 bases | Strand | Plus strand |
| Position | 17 : 57255851 - 57684689 | RNA binding domain | RRM_1 | ||
| Summary | This gene encodes an RNA-binding protein that is a member of the Musashi protein family. The encoded protein is transcriptional regulator that targets genes involved in development and cell cycle regulation. Mutations in this gene are associated with poor prognosis in certain types of cancers. This gene has also been shown to be rearranged in certain cancer cells. [provided by RefSeq, Apr 2016] | ||||
RNA binding domains (RBDs)
| Protein | Domain | Pfam ID | E-value | Domain number |
|---|---|---|---|---|
| ENSP00000313616 | RRM_1 | PF00076 | 4e-34 | 1 |
| ENSP00000313616 | RRM_1 | PF00076 | 4e-34 | 2 |
| ENSP00000414671 | RRM_1 | PF00076 | 7.8e-34 | 1 |
| ENSP00000414671 | RRM_1 | PF00076 | 7.8e-34 | 2 |
| ENSP00000501595 | RRM_1 | PF00076 | 9.3e-34 | 1 |
| ENSP00000501595 | RRM_1 | PF00076 | 9.3e-34 | 2 |
| ENSP00000284073 | RRM_1 | PF00076 | 9.4e-34 | 1 |
| ENSP00000284073 | RRM_1 | PF00076 | 9.4e-34 | 2 |
| ENSP00000502137 | RRM_1 | PF00076 | 9.4e-34 | 1 |
| ENSP00000502137 | RRM_1 | PF00076 | 9.4e-34 | 2 |
| ENSP00000462914 | RRM_1 | PF00076 | 1.2e-22 | 1 |
| ENSP00000502813 | RRM_1 | PF00076 | 1.3e-17 | 1 |
| ENSP00000501981 | RRM_1 | PF00076 | 2.2e-17 | 1 |
| ENSP00000463227 | RRM_1 | PF00076 | 2.4e-17 | 1 |
| ENSP00000462264 | RRM_1 | PF00076 | 1.5e-14 | 1 |
| ENSP00000392607 | RRM_1 | PF00076 | 4.3e-14 | 1 |
| ENSP00000502212 | RRM_1 | PF00076 | 1.3e-05 | 1 |
RNA binding proteomes (RBPomes)
| Pubmed ID | Full Name | Cell | Author | Time | Doi |
|---|
Literatures on RNA binding capacity
| Pubmed ID | Title | Author | Time | Journal |
|---|---|---|---|---|
| 31952541 | Musashi2 promotes EGF-induced EMT in pancreatic cancer via ZEB1-ERK/MAPK signaling. | Weiwei Sheng | 2020-01-17 | Journal of experimental & clinical cancer research : CR |
| 31217428 | Small-molecule targeting of MUSASHI RNA-binding activity in acute myeloid leukemia. | Gerard Minuesa | 2019-06-19 | Nature communications |
| 27593929 | Musashi RNA-binding protein 2 regulates estrogen receptor 1 function in breast cancer. | M-H Kang | 2017-03-23 | Oncogene |
| 33260776 | Impact of AHR Ligand TCDD on Human Embryonic Stem Cells and Early Differentiation. | Indrek Teino | 2020-11-28 | International journal of molecular sciences |
| 36153373 | The Musashi proteins direct post-transcriptional control of protein expression and alternate exon splicing in vertebrate photoreceptors. | Fatimah Matalkah | 2022-09-24 | Communications biology |
| 31888685 | Musashi2 contributes to the maintenance of CD44v6+ liver cancer stem cells via notch1 signaling pathway. | Xiju Wang | 2019-12-30 | Journal of experimental & clinical cancer research : CR |
| 37076565 | Convergent genomic diversity and novel BCAA metabolism in intrahepatic cholangiocarcinoma. | Akihiro Kitagawa | 2023-06-01 | British journal of cancer |
| 34237057 | Prognostic role and biologic features of Musashi-2 expression in colon polyps and during colorectal cancer progression. | Leonid Kharin | 2021-01-01 | PloS one |
| 37117191 | MSI2 promotes translation of multiple IRES-containing oncogenes and virus to induce self-renewal of tumor initiating stem-like cells. | Da-Wei Yeh | 2023-04-28 | Cell death discovery |
| 27092875 | Musashi2 promotes the development and progression of pancreatic cancer by down-regulating Numb protein. | Weiwei Sheng | 2017-02-28 | Oncotarget |
| 27307150 | Cellular and Molecular Networks in Chronic Myeloid Leukemia: The Leukemic Stem, Progenitor and Stromal Cell Interplay. | Danilo Perrotti | 2017-01-01 | Current drug targets |
| 30034243 | MSI2 knockdown represses extrahepatic cholangiocarcinoma growth and invasion by inhibiting epithelial-mesenchymal transition. | Feihu Hu | 2018-01-01 | OncoTargets and therapy |
| 30836959 | High-resolution population structure and runs of homozygosity reveal the genetic architecture of complex traits in the Lipizzan horse. | Gertrud Grilz-Seger | 2019-03-05 | BMC genomics |
| 28107692 | RNA binding protein MSI2 positively regulates FLT3 expression in myeloid leukemia. | Ayuna Hattori | 2017-03-01 | Leukemia research |
| 34107032 | RNA pull-down confocal nanoscanning (RP-CONA) detects quercetin as pri-miR-7/HuR interaction inhibitor that decreases α-synuclein levels. | Siran Zhu | 2021-06-21 | Nucleic acids research |
| 28514443 | Cancer progression by reprogrammed BCAA metabolism in myeloid leukaemia. | Ayuna Hattori | 2017-05-25 | Nature |
| 37149661 | MicroRNA-143 acts as a tumor suppressor through Musashi-2/DLL1/Notch1 and Musashi-2/Snail1/MMPs axes in acute myeloid leukemia. | Fanfan Li | 2023-05-06 | Journal of translational medicine |
| 29739919 | Identification of Core Genes and Key Pathways via Integrated Analysis of Gene Expression and DNA Methylation Profiles in Bladder Cancer. | Yongzhen Zhang | 2018-05-09 | Medical science monitor : international medical journal of experimental and clinical research |
| 33723247 | Musashi-2 (MSI2) regulates epidermal growth factor receptor (EGFR) expression and response to EGFR inhibitors in EGFR-mutated non-small cell lung cancer (NSCLC). | Petr Makhov | 2021-03-15 | Oncogenesis |
| 33504942 | Musashi 2 influences chronic lymphocytic leukemia cell survival and growth making it a potential therapeutic target. | Florencia Palacios | 2021-04-01 | Leukemia |
| 31878037 | Immunohistochemical Analysis Revealed a Correlation between Musashi-2 and Cyclin-D1 Expression in Patients with Oral Squamous Cells Carcinoma. | Giuseppe Troiano | 2019-12-23 | International journal of molecular sciences |
| 35528200 | Integrative genome-wide analysis reveals EIF3A as a key downstream regulator of translational repressor protein Musashi 2 (MSI2). | Shilpita Karmakar | 2022-06-01 | NAR cancer |
| 30719075 | Targeting liver cancer stem cells for the treatment of hepatocellular carcinoma. | Na Li | 2019-01-01 | Therapeutic advances in gastroenterology |
| 35153752 | Small Molecule Palmatine Targeting Musashi-2 in Colorectal Cancer. | Xue Zhang | 2021-01-01 | Frontiers in pharmacology |
| 32606289 | Knockdown of MSI2 inhibits metastasis by interacting with caveolin-1 and inhibiting its ubiquitylation in human NF1-MPNST cells. | Kang Yang | 2020-06-30 | Cell death & disease |
| 37173995 | Regulation of VEGFR2 and AKT Signaling by Musashi-2 in Lung Cancer. | Igor Bychkov | 2023-04-28 | Cancers |
| 37034813 | Regulation of VEGFR2 and AKT signaling by Musashi-2 in lung cancer. | Igor Bychkov | 2023-03-31 | bioRxiv : the preprint server for biology |
| 29073938 | Musashi-2, a novel oncoprotein promoting cervical cancer cell growth and invasion, is negatively regulated by p53-induced miR-143 and miR-107 activation. | Peixin Dong | 2017-10-26 | Journal of experimental & clinical cancer research : CR |
| 33553241 | RNA-Binding Protein MSI2 Binds to miR-301a-3p and Facilitates Its Distribution in Mitochondria of Endothelial Cells. | Qian Qian Guo | 2020-01-01 | Frontiers in molecular biosciences |
Transcripts
| Name | Transcript ID | bp | Protein | Translation ID |
|---|---|---|---|---|
| MSI2-203 | ENST00000416426 | 1714 | 324aa | ENSP00000414671 |
| MSI2-201 | ENST00000284073 | 6379 | 328aa | ENSP00000284073 |
| MSI2-224 | ENST00000674964 | 6301 | 328aa | ENSP00000502137 |
| MSI2-214 | ENST00000581776 | 493 | 99aa | ENSP00000463227 |
| MSI2-213 | ENST00000581523 | 534 | No protein | - |
| MSI2-223 | ENST00000674961 | 2124 | 107aa | ENSP00000501981 |
| MSI2-215 | ENST00000582453 | 496 | No protein | - |
| MSI2-227 | ENST00000675656 | 2247 | 341aa | ENSP00000501595 |
| MSI2-202 | ENST00000322684 | 1860 | 251aa | ENSP00000313616 |
| MSI2-212 | ENST00000579590 | 452 | 140aa | ENSP00000462914 |
| MSI2-211 | ENST00000579531 | 472 | No protein | - |
| MSI2-230 | ENST00000676105 | 246 | 31aa | ENSP00000502565 |
| MSI2-229 | ENST00000676073 | 425 | 89aa | ENSP00000502813 |
| MSI2-219 | ENST00000584476 | 465 | No protein | - |
| MSI2-209 | ENST00000579483 | 5141 | No protein | - |
| MSI2-228 | ENST00000675822 | 331 | 34aa | ENSP00000502227 |
| MSI2-204 | ENST00000442934 | 1624 | 267aa | ENSP00000392607 |
| MSI2-206 | ENST00000579180 | 1635 | 151aa | ENSP00000462264 |
| MSI2-207 | ENST00000579205 | 500 | No protein | - |
| MSI2-210 | ENST00000579505 | 577 | No protein | - |
| MSI2-216 | ENST00000582772 | 398 | No protein | - |
| MSI2-222 | ENST00000674898 | 1921 | 139aa | ENSP00000502212 |
| MSI2-231 | ENST00000676345 | 119 | No protein | - |
| MSI2-221 | ENST00000674574 | 274 | 22aa | ENSP00000502773 |
| MSI2-205 | ENST00000577241 | 710 | No protein | - |
| MSI2-217 | ENST00000583705 | 7612 | No protein | - |
| MSI2-218 | ENST00000583821 | 548 | No protein | - |
| MSI2-225 | ENST00000675127 | 341 | 73aa | ENSP00000502546 |
| MSI2-226 | ENST00000675379 | 5204 | 41aa | ENSP00000502532 |
| MSI2-208 | ENST00000579466 | 186 | No protein | - |
| MSI2-220 | ENST00000674522 | 5296 | 65aa | ENSP00000501836 |
Phenotypes
| ensgID | Trait | pValue | Pubmed ID |
|---|---|---|---|
| 18971 | Iron | 8.946e-005 | - |
| 19464 | Electrocardiography | 1.111e-005 | - |
| 26177 | Arteries | 9.3240000E-005 | - |
| 26317 | Arteries | 5.3060000E-005 | - |
| 26346 | Arteries | 5.6520000E-005 | - |
| 26348 | Arteries | 4.5840000E-005 | - |
| 30374 | Potassium | 3.7770000E-005 | - |
| 41879 | Platelet Function Tests | 5.8976601E-006 | - |
| 43030 | Platelet Function Tests | 1.6541500E-005 | - |
| 46803 | Platelet Function Tests | 1.4219000E-005 | - |
| 47046 | Platelet Function Tests | 8.3463000E-006 | - |
| 49080 | Child Development Disorders, Pervasive | 3.0537600E-005 | - |
| 52210 | Child Development Disorders, Pervasive | 5.4000000E-005 | - |
| 74330 | Schizophrenia | 2E-6 | 26198764 |
| 92188 | Vascular Calcification | 8E-6 | 23870195 |
GWAS
| ensgID | SNP | Chromosome | Position | Trait | PubmedID | Or or BEAT | EFO ID |
|---|
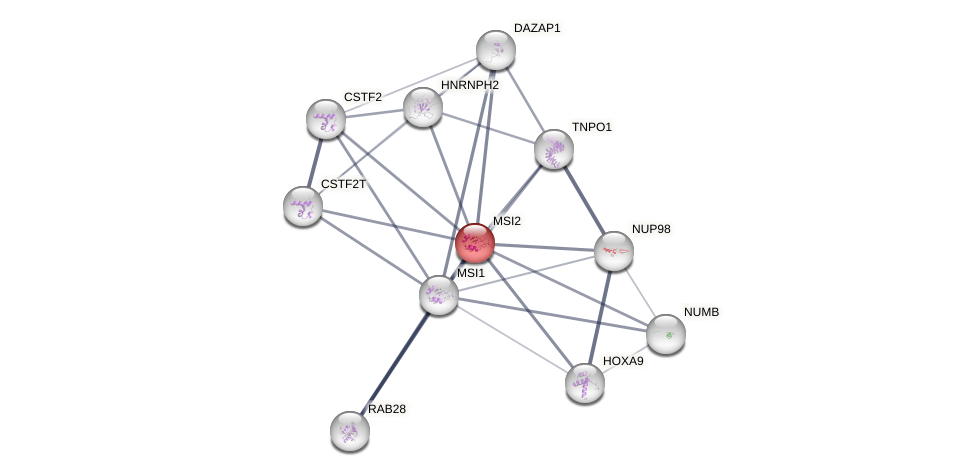
Protein-Protein Interaction (PPI)
Paralogs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| ENSG00000153944 | homo_sapiens | ENSP00000295971 | 15.5143 | 8.76897 | homo_sapiens | ENSP00000284073 | 28.0488 | 15.8537 |
| ENSG00000153944 | homo_sapiens | ENSP00000304139 | 26.5306 | 18.8776 | homo_sapiens | ENSP00000284073 | 31.7073 | 22.561 |
| ENSG00000153944 | homo_sapiens | ENSP00000417451 | 8.52858 | 5.62324 | homo_sapiens | ENSP00000284073 | 27.7439 | 18.2927 |
| ENSG00000153944 | homo_sapiens | ENSP00000478691 | 48.0938 | 31.9648 | homo_sapiens | ENSP00000284073 | 50 | 33.2317 |
| ENSG00000153944 | homo_sapiens | ENSP00000445366 | 20.1133 | 11.3314 | homo_sapiens | ENSP00000284073 | 21.6463 | 12.1951 |
| ENSG00000153944 | homo_sapiens | ENSP00000341826 | 45.9677 | 30.3763 | homo_sapiens | ENSP00000284073 | 52.1341 | 34.4512 |
| ENSG00000153944 | homo_sapiens | ENSP00000350090 | 52.8125 | 35.625 | homo_sapiens | ENSP00000284073 | 51.5244 | 34.7561 |
| ENSG00000153944 | homo_sapiens | ENSP00000493805 | 53.125 | 34.6875 | homo_sapiens | ENSP00000284073 | 51.8293 | 33.8415 |
| ENSG00000153944 | homo_sapiens | ENSP00000376309 | 44.1799 | 31.4815 | homo_sapiens | ENSP00000284073 | 50.9146 | 36.2805 |
| ENSG00000153944 | homo_sapiens | ENSP00000310471 | 26.1838 | 14.4847 | homo_sapiens | ENSP00000284073 | 28.6585 | 15.8537 |
| ENSG00000153944 | homo_sapiens | ENSP00000309166 | 25.2747 | 14.011 | homo_sapiens | ENSP00000284073 | 28.0488 | 15.5488 |
| ENSG00000153944 | homo_sapiens | ENSP00000498248 | 30 | 20 | homo_sapiens | ENSP00000284073 | 35.6707 | 23.7805 |
| ENSG00000153944 | homo_sapiens | ENSP00000359645 | 30.4348 | 20.4604 | homo_sapiens | ENSP00000284073 | 36.2805 | 24.3902 |
| ENSG00000153944 | homo_sapiens | ENSP00000401371 | 27.7202 | 15.8031 | homo_sapiens | ENSP00000284073 | 32.622 | 18.5976 |
| ENSG00000153944 | homo_sapiens | ENSP00000257552 | 75.1381 | 68.5083 | homo_sapiens | ENSP00000284073 | 82.9268 | 75.6098 |
| ENSG00000153944 | homo_sapiens | ENSP00000281722 | 18.5741 | 10.5066 | homo_sapiens | ENSP00000284073 | 30.1829 | 17.0732 |
| ENSG00000153944 | homo_sapiens | ENSP00000352956 | 17.0843 | 10.9339 | homo_sapiens | ENSP00000284073 | 22.8659 | 14.6341 |
| ENSG00000153944 | homo_sapiens | ENSP00000362936 | 25.0871 | 13.2404 | homo_sapiens | ENSP00000284073 | 21.9512 | 11.5854 |
| ENSG00000153944 | homo_sapiens | ENSP00000313199 | 38.0282 | 29.0141 | homo_sapiens | ENSP00000284073 | 41.1585 | 31.4024 |
| ENSG00000153944 | homo_sapiens | ENSP00000253363 | 16.6038 | 10.3774 | homo_sapiens | ENSP00000284073 | 26.8293 | 16.7683 |
| ENSG00000153944 | homo_sapiens | ENSP00000363109 | 17.5768 | 10.7509 | homo_sapiens | ENSP00000284073 | 31.4024 | 19.2073 |
| ENSG00000153944 | homo_sapiens | ENSP00000316042 | 49.8361 | 36.7213 | homo_sapiens | ENSP00000284073 | 46.3415 | 34.1463 |
| ENSG00000153944 | homo_sapiens | ENSP00000318195 | 13.662 | 6.76056 | homo_sapiens | ENSP00000284073 | 29.5732 | 14.6341 |
| ENSG00000153944 | homo_sapiens | ENSP00000394902 | 27.4667 | 15.7333 | homo_sapiens | ENSP00000284073 | 31.4024 | 17.9878 |
| ENSG00000153944 | homo_sapiens | ENSP00000351108 | 42.1687 | 31.9277 | homo_sapiens | ENSP00000284073 | 42.6829 | 32.3171 |
| ENSG00000153944 | homo_sapiens | ENSP00000261741 | 11.1458 | 6.97917 | homo_sapiens | ENSP00000284073 | 32.622 | 20.4268 |
| ENSG00000153944 | homo_sapiens | ENSP00000233078 | 28.5012 | 19.656 | homo_sapiens | ENSP00000284073 | 35.3659 | 24.3902 |
| ENSG00000153944 | homo_sapiens | ENSP00000372127 | 19.5565 | 11.4919 | homo_sapiens | ENSP00000284073 | 29.5732 | 17.378 |
| ENSG00000153944 | homo_sapiens | ENSP00000372105 | 19.5565 | 11.4919 | homo_sapiens | ENSP00000284073 | 29.5732 | 17.378 |
| ENSG00000153944 | homo_sapiens | ENSP00000372484 | 19.7581 | 11.6935 | homo_sapiens | ENSP00000284073 | 29.878 | 17.6829 |
| ENSG00000153944 | homo_sapiens | ENSP00000372154 | 19.5565 | 11.4919 | homo_sapiens | ENSP00000284073 | 29.5732 | 17.378 |
| ENSG00000153944 | homo_sapiens | ENSP00000250831 | 19.9597 | 11.8952 | homo_sapiens | ENSP00000284073 | 30.1829 | 17.9878 |
| ENSG00000153944 | homo_sapiens | ENSP00000307155 | 19.9597 | 11.8952 | homo_sapiens | ENSP00000284073 | 30.1829 | 17.9878 |
| ENSG00000153944 | homo_sapiens | ENSP00000295470 | 31.6667 | 25 | homo_sapiens | ENSP00000284073 | 40.5488 | 32.0122 |
| ENSG00000153944 | homo_sapiens | ENSP00000466025 | 28.6195 | 19.8653 | homo_sapiens | ENSP00000284073 | 25.9146 | 17.9878 |
| ENSG00000153944 | homo_sapiens | ENSP00000365950 | 47.7707 | 35.6688 | homo_sapiens | ENSP00000284073 | 22.8659 | 17.0732 |
Orthologs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.3293 | 89.3293 | nomascus_leucogenys | ENSNLEP00000009843 | 94.2122 | 94.2122 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | pan_paniscus | ENSPPAP00000029419 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 92.9878 | pongo_abelii | ENSPPYP00000027665 | 94.1358 | 94.1358 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | pan_troglodytes | ENSPTRP00000016029 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | gorilla_gorilla | ENSGGOP00000012862 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.2073 | 92.0732 | ornithorhynchus_anatinus | ENSOANP00000053353 | 55.7762 | 54.5126 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.1707 | 97.8659 | sarcophilus_harrisii | ENSSHAP00000039221 | 88.2192 | 87.9452 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 65.2439 | 65.2439 | notamacropus_eugenii | ENSMEUP00000003851 | 80.4511 | 80.4511 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 60.061 | 59.7561 | choloepus_hoffmanni | ENSCHOP00000007666 | 70.3571 | 70 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.8049 | 87.8049 | dasypus_novemcinctus | ENSDNOP00000011239 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 66.1585 | 65.8537 | erinaceus_europaeus | ENSEEUP00000007115 | 87.1486 | 86.747 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 39.6341 | 39.3293 | echinops_telfairi | ENSETEP00000010094 | 54.6218 | 54.2017 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | callithrix_jacchus | ENSCJAP00000025750 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.7805 | 98.7805 | cercocebus_atys | ENSCATP00000036127 | 98.7805 | 98.7805 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 86.5854 | 84.7561 | macaca_mulatta | ENSMMUP00000055809 | 83.5294 | 81.7647 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | macaca_nemestrina | ENSMNEP00000045534 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | papio_anubis | ENSPANP00000020344 | 94.7977 | 94.7977 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | mandrillus_leucophaeus | ENSMLEP00000019380 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 65.8537 | 65.8537 | tursiops_truncatus | ENSTTRP00000002052 | 90.7563 | 90.7563 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 91.7683 | vulpes_vulpes | ENSVVUP00000016032 | 67.0354 | 66.5929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | mus_spretus | MGP_SPRETEiJ_P0030545 | 94.5087 | 94.5087 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 96.9512 | 96.6463 | equus_asinus | ENSEASP00005036209 | 83.4646 | 83.2021 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.4756 | 98.4756 | capra_hircus | ENSCHIP00000016257 | 98.4756 | 98.4756 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 50 | 50 | loxodonta_africana | ENSLAFP00000003143 | 90.1099 | 90.1099 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 61.2805 | 59.4512 | panthera_leo | ENSPLOP00000024976 | 81.7073 | 79.2683 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 40.8537 | 39.3293 | ailuropoda_melanoleuca | ENSAMEP00000043091 | 76.5714 | 73.7143 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 86.2805 | 83.2317 | bos_indicus_hybrid | ENSBIXP00005024733 | 70.0495 | 67.5743 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | oryctolagus_cuniculus | ENSOCUP00000008989 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | equus_caballus | ENSECAP00000033438 | 86.0892 | 86.0892 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 81.4024 | 78.6585 | canis_lupus_familiaris | ENSCAFP00845023022 | 70.0787 | 67.7165 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.5976 | 93.5976 | panthera_pardus | ENSPPRP00000006373 | 93.5976 | 93.5976 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.9268 | 82.0122 | marmota_marmota_marmota | ENSMMMP00000023688 | 91.5825 | 90.5724 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | balaenoptera_musculus | ENSBMSP00010028012 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.6219 | 82.0122 | cavia_porcellus | ENSCPOP00000027036 | 95.7597 | 95.053 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 55.1829 | 51.8293 | felis_catus | ENSFCAP00000028027 | 69.084 | 64.8855 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.4268 | 95.1219 | ursus_americanus | ENSUAMP00000000698 | 96.6049 | 96.2963 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 56.4024 | 56.4024 | monodelphis_domestica | ENSMODP00000059304 | 86.0465 | 86.0465 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 44.5122 | 43.5976 | bison_bison_bison | ENSBBBP00000002067 | 97.3333 | 95.3333 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 96.9512 | 96.9512 | monodon_monoceros | ENSMMNP00015021334 | 88.3333 | 88.3333 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.7317 | 95.7317 | sus_scrofa | ENSSSCP00000043030 | 92.0821 | 92.0821 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 97.561 | 97.2561 | microcebus_murinus | ENSMICP00000048820 | 88.1543 | 87.8788 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 84.4512 | 81.4024 | microcebus_murinus | ENSMICP00000035082 | 94.2177 | 90.8163 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | octodon_degus | ENSODEP00000016268 | 94.7977 | 94.7977 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 53.0488 | 51.8293 | jaculus_jaculus | ENSJJAP00000003258 | 87.4372 | 85.4271 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 34.4512 | 34.4512 | jaculus_jaculus | ENSJJAP00000004511 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 92.9878 | ovis_aries_rambouillet | ENSOARP00020028592 | 94.1358 | 94.1358 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 75.3049 | 75 | ursus_maritimus | ENSUMAP00000000373 | 96.1089 | 95.7198 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 45.122 | 44.8171 | ochotona_princeps | ENSOPRP00000000801 | 62.7119 | 62.2881 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | delphinapterus_leucas | ENSDLEP00000008988 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 22.561 | 20.122 | physeter_catodon | ENSPCTP00005017944 | 41.573 | 37.0787 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | mus_caroli | MGP_CAROLIEiJ_P0029133 | 94.5087 | 94.5087 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | mus_musculus | ENSMUSP00000090470 | 94.5087 | 94.5087 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 34.4512 | 34.4512 | dipodomys_ordii | ENSDORP00000023713 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.4268 | 95.1219 | mustela_putorius_furo | ENSMPUP00000015267 | 96.6049 | 96.2963 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | aotus_nancymaae | ENSANAP00000041004 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.2927 | 93.2927 | saimiri_boliviensis_boliviensis | ENSSBOP00000019694 | 94.4444 | 94.4444 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 50.9146 | 50.9146 | vicugna_pacos | ENSVPAP00000006353 | 56.9966 | 56.9966 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | chlorocebus_sabaeus | ENSCSAP00000001941 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 66.7683 | 64.6341 | tupaia_belangeri | ENSTBEP00000001089 | 67.8019 | 65.6347 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | mesocricetus_auratus | ENSMAUP00000003504 | 94.2363 | 94.2363 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | phocoena_sinus | ENSPSNP00000019277 | 94.7977 | 94.7977 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.3293 | 89.3293 | camelus_dromedarius | ENSCDRP00005032533 | 94.2122 | 94.2122 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 57.3171 | 55.7927 | carlito_syrichta | ENSTSYP00000010859 | 84.3049 | 82.0628 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 62.8049 | 60.3659 | panthera_tigris_altaica | ENSPTIP00000026464 | 84.7737 | 81.4815 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 96.6463 | 96.3415 | rattus_norvegicus | ENSRNOP00000096507 | 91.6185 | 91.3295 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 44.2073 | 43.5976 | prolemur_simus | ENSPSMP00000014134 | 96.0265 | 94.702 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | microtus_ochrogaster | ENSMOCP00000006239 | 94.5087 | 94.5087 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.5976 | 93.2927 | peromyscus_maniculatus_bairdii | ENSPEMP00000025331 | 88.7283 | 88.4393 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 44.5122 | 43.5976 | bos_mutus | ENSBMUP00000000017 | 97.3333 | 95.3333 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.7805 | 98.4756 | mus_spicilegus | ENSMSIP00000027026 | 93.6416 | 93.3526 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | cricetulus_griseus_chok1gshd | ENSCGRP00001011768 | 99.3921 | 99.3921 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 81.0976 | 81.0976 | pteropus_vampyrus | ENSPVAP00000016085 | 90.785 | 90.785 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 94.8171 | myotis_lucifugus | ENSMLUP00000007176 | 96.2963 | 95.9877 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.1707 | 97.8659 | vombatus_ursinus | ENSVURP00010017701 | 93.0636 | 92.7746 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 92.9878 | rhinolophus_ferrumequinum | ENSRFEP00010024966 | 94.4272 | 94.4272 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.5976 | 93.5976 | neovison_vison | ENSNVIP00000015876 | 93.5976 | 93.5976 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 92.3781 | bos_taurus | ENSBTAP00000059519 | 84.2541 | 83.7017 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 92.0732 | heterocephalus_glaber_female | ENSHGLP00000055602 | 88.1503 | 87.2832 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 25.9146 | 25.3049 | rhinopithecus_bieti | ENSRBIP00000015975 | 91.3979 | 89.2473 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.6341 | 89.6341 | canis_lupus_dingo | ENSCAFP00020032985 | 74.8092 | 74.8092 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | sciurus_vulgaris | ENSSVLP00005012124 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.1707 | 97.8659 | phascolarctos_cinereus | ENSPCIP00000007136 | 88.2192 | 87.9452 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 51.2195 | 51.2195 | procavia_capensis | ENSPCAP00000009813 | 70.5882 | 70.5882 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | nannospalax_galili | ENSNGAP00000017354 | 94.5087 | 94.5087 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 84.7561 | 83.2317 | bos_grunniens | ENSBGRP00000025839 | 80.5797 | 79.1304 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | chinchilla_lanigera | ENSCLAP00000020905 | 94.7977 | 94.7977 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.3293 | 89.3293 | cervus_hanglu_yarkandensis | ENSCHYP00000021527 | 90.4321 | 90.4321 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.7195 | 88.7195 | catagonus_wagneri | ENSCWAP00000010810 | 94.1748 | 94.1748 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.6219 | 82.0122 | ictidomys_tridecemlineatus | ENSSTOP00000007055 | 95.7597 | 95.053 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.3293 | 89.0244 | otolemur_garnettii | ENSOGAP00000005024 | 95.4397 | 95.114 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 100 | 100 | cebus_imitator | ENSCCAP00000031844 | 100 | 100 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.9268 | 82.0122 | urocitellus_parryii | ENSUPAP00010018071 | 86.3492 | 85.3968 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.7317 | 95.4268 | moschus_moschiferus | ENSMMSP00000018175 | 95.7317 | 95.4268 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 53.9634 | 53.9634 | rhinopithecus_roxellana | ENSRROP00000015182 | 89.8477 | 89.8477 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 99.6951 | 99.6951 | mus_pahari | MGP_PahariEiJ_P0035846 | 94.5087 | 94.5087 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 50 | 49.3902 | propithecus_coquereli | ENSPCOP00000024750 | 81.1881 | 80.198 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 59.1463 | 48.1707 | caenorhabditis_elegans | R10E9.1.1 | 60.625 | 49.375 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 61.5854 | 46.3415 | drosophila_melanogaster | FBpp0305820 | 31.8612 | 23.9748 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 68.2927 | 57.622 | drosophila_melanogaster | FBpp0112339 | 44.8898 | 37.8758 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 41.1585 | 35.9756 | ciona_intestinalis | ENSCINP00000031642 | 60.8108 | 53.1532 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 70.7317 | 64.3293 | eptatretus_burgeri | ENSEBUP00000010477 | 67.8363 | 61.6959 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 80.4878 | 78.0488 | callorhinchus_milii | ENSCMIP00000022186 | 75.6447 | 73.3524 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 26.8293 | 26.2195 | latimeria_chalumnae | ENSLACP00000019089 | 87.1287 | 85.1485 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 96.0366 | 95.4268 | lepisosteus_oculatus | ENSLOCP00000005599 | 81.3953 | 80.8786 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.1951 | 85.3659 | clupea_harengus | ENSCHAP00000017385 | 69.4175 | 67.9612 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.0732 | 91.1585 | clupea_harengus | ENSCHAP00000056546 | 71.9048 | 71.1905 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | danio_rerio | ENSDARP00000123512 | 80.4124 | 78.3505 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.7317 | 94.8171 | danio_rerio | ENSDARP00000090967 | 76.9608 | 76.2255 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 88.4146 | carassius_auratus | ENSCARP00000060517 | 65.5702 | 63.5965 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.4146 | 84.1463 | carassius_auratus | ENSCARP00000061674 | 70.2179 | 66.8281 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 23.1707 | 22.561 | carassius_auratus | ENSCARP00000069386 | 89.4118 | 87.0588 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.1098 | 86.5854 | carassius_auratus | ENSCARP00000070803 | 73.7245 | 72.449 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 65.8537 | 63.4146 | carassius_auratus | ENSCARP00000115948 | 65.0602 | 62.6506 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 92.0732 | astyanax_mexicanus | ENSAMXP00000046855 | 78.4062 | 77.635 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.7317 | 93.9024 | astyanax_mexicanus | ENSAMXP00000049152 | 80.7198 | 79.1774 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.5976 | 91.7683 | ictalurus_punctatus | ENSIPUP00000005377 | 75.9901 | 74.505 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.9024 | 90.5488 | ictalurus_punctatus | ENSIPUP00000031344 | 69.0583 | 66.5919 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.8049 | 86.2805 | esox_lucius | ENSELUP00000016414 | 62.3377 | 61.2554 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.0732 | 90.5488 | esox_lucius | ENSELUP00000033503 | 70.0696 | 68.9095 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.2439 | 87.1951 | electrophorus_electricus | ENSEEEP00000020227 | 72.1951 | 69.7561 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.8171 | 92.3781 | electrophorus_electricus | ENSEEEP00000025860 | 70.5215 | 68.7075 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 86.5854 | 84.7561 | oncorhynchus_kisutch | ENSOKIP00005018250 | 67.1395 | 65.721 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.0732 | 90.5488 | oncorhynchus_kisutch | ENSOKIP00005106119 | 69.4253 | 68.2759 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.4146 | 85.9756 | oncorhynchus_kisutch | ENSOKIP00005041290 | 64.3016 | 62.5277 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.7683 | 90.5488 | oncorhynchus_kisutch | ENSOKIP00005112624 | 73.9558 | 72.973 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.5122 | 93.5976 | oncorhynchus_mykiss | ENSOMYP00000120366 | 76.3547 | 75.6158 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.6219 | 77.7439 | oncorhynchus_mykiss | ENSOMYP00000076065 | 60.8989 | 57.3034 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 86.5854 | 84.7561 | oncorhynchus_mykiss | ENSOMYP00000089810 | 67.1395 | 65.721 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.0732 | 90.5488 | oncorhynchus_mykiss | ENSOMYP00000130064 | 69.4253 | 68.2759 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.7683 | 90.5488 | salmo_salar | ENSSSAP00000070529 | 71.327 | 70.3792 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.7683 | 89.3293 | salmo_salar | ENSSSAP00000122764 | 66.1538 | 64.3956 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.2073 | 93.2927 | salmo_salar | ENSSSAP00000100919 | 72.5352 | 71.831 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 89.3293 | salmo_salar | ENSSSAP00000133751 | 71.5311 | 70.0957 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.8537 | 89.0244 | salmo_trutta | ENSSTUP00000086701 | 68.1922 | 66.8192 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.7683 | 89.6341 | salmo_trutta | ENSSTUP00000026899 | 69.0367 | 67.4312 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.2073 | 93.2927 | salmo_trutta | ENSSTUP00000011738 | 70.5479 | 69.863 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 89.939 | salmo_trutta | ENSSTUP00000011041 | 73.2843 | 72.3039 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 19.8171 | 16.4634 | gadus_morhua | ENSGMOP00000025765 | 59.0909 | 49.0909 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.3293 | fundulus_heteroclitus | ENSFHEP00000002093 | 74.2647 | 71.8137 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | poecilia_reticulata | ENSPREP00000001190 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | xiphophorus_maculatus | ENSXMAP00000027105 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.6341 | oryzias_latipes | ENSORLP00000043770 | 77.892 | 75.5784 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.8049 | 85.061 | cyclopterus_lumpus | ENSCLMP00005051044 | 74.2268 | 71.9072 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 81.7073 | 77.7439 | oreochromis_niloticus | ENSONIP00000072063 | 55.144 | 52.4691 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | haplochromis_burtoni | ENSHBUP00000033716 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | astatotilapia_calliptera | ENSACLP00000011353 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.4146 | 85.061 | sparus_aurata | ENSSAUP00010026690 | 69.7115 | 67.0673 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | lates_calcarifer | ENSLCAP00010047697 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.6219 | 78.6585 | lates_calcarifer | ENSLCAP00010013416 | 57.4153 | 54.661 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 97.561 | 96.6463 | xenopus_tropicalis | ENSXETP00000090211 | 77.1084 | 76.3855 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 81.7073 | 81.7073 | chrysemys_picta_bellii | ENSCPBP00000013383 | 70.3412 | 70.3412 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 53.0488 | 52.439 | sphenodon_punctatus | ENSSPUP00000001142 | 85.2941 | 84.3137 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 48.1707 | 46.6463 | sphenodon_punctatus | ENSSPUP00000001151 | 93.4911 | 90.5325 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 90.8537 | crocodylus_porosus | ENSCPRP00005014494 | 75.1861 | 73.9454 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 21.3415 | 19.5122 | laticauda_laticaudata | ENSLLTP00000002739 | 63.6364 | 58.1818 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 26.5244 | 25 | notechis_scutatus | ENSNSUP00000016850 | 91.5789 | 86.3158 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 54.5732 | 54.2683 | notechis_scutatus | ENSNSUP00000016851 | 87.7451 | 87.2549 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 97.8659 | 96.6463 | pseudonaja_textilis | ENSPTXP00000016303 | 79.2593 | 78.2716 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 22.2561 | 19.8171 | anas_platyrhynchos_platyrhynchos | ENSAPLP00000032048 | 56.1538 | 50 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 93.9024 | 93.2927 | gallus_gallus | ENSGALP00010044041 | 86.2745 | 85.7143 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 21.9512 | 19.8171 | meleagris_gallopavo | ENSMGAP00000008484 | 57.1429 | 51.5873 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 26.2195 | 24.3902 | serinus_canaria | ENSSCAP00000015965 | 76.1062 | 70.7965 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 64.6341 | 64.3293 | parus_major | ENSPMJP00000027286 | 94.6429 | 94.1964 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 84.1463 | 82.6219 | anolis_carolinensis | ENSACAP00000038428 | 70.4082 | 69.1327 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | neolamprologus_brichardi | ENSNBRP00000018941 | 80.4124 | 78.0928 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 96.9512 | 94.8171 | naja_naja | ENSNNAP00000026677 | 67.6596 | 66.1702 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.5122 | 93.5976 | podarcis_muralis | ENSPMRP00000030102 | 94.8012 | 93.8838 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 43.9024 | 43.2927 | geospiza_fortis | ENSGFOP00000010293 | 95.3642 | 94.0397 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 92.3781 | taeniopygia_guttata | ENSTGUP00000008709 | 93.5185 | 93.5185 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 80.4878 | 78.6585 | erpetoichthys_calabaricus | ENSECRP00000009720 | 70.2128 | 68.617 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.8171 | 91.7683 | kryptolebias_marmoratus | ENSKMAP00000012709 | 80.1546 | 77.5773 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 89.6341 | salvator_merianae | ENSSMRP00000016991 | 73.8272 | 72.5926 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.0122 | 78.6585 | oryzias_javanicus | ENSOJAP00000026474 | 63.5934 | 60.9929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.5488 | 87.8049 | amphilophus_citrinellus | ENSACIP00000012180 | 78.7798 | 76.3926 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.2439 | 88.4146 | dicentrarchus_labrax | ENSDLAP00005083160 | 69.9764 | 68.5579 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.6341 | dicentrarchus_labrax | ENSDLAP00005050101 | 74.2647 | 72.0588 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 81.0976 | 77.7439 | amphiprion_percula | ENSAPEP00000010762 | 60.181 | 57.6923 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.6341 | oryzias_sinensis | ENSOSIP00000039945 | 74.2647 | 72.0588 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.4756 | 98.1707 | pelodiscus_sinensis | ENSPSIP00000009465 | 79.7531 | 79.5062 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.4756 | 98.1707 | gopherus_evgoodei | ENSGEVP00005025425 | 79.5566 | 79.3103 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.6829 | 91.4634 | chelonoidis_abingdonii | ENSCABP00000023523 | 92.9664 | 91.7431 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | seriola_dumerili | ENSSDUP00000019978 | 76.6585 | 74.6929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.4146 | 86.5854 | seriola_dumerili | ENSSDUP00000030516 | 87.0871 | 85.2853 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.6341 | cottoperca_gobio | ENSCGOP00000053389 | 74.2647 | 72.0588 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 86.5854 | 85.061 | cottoperca_gobio | ENSCGOP00000026623 | 69.6078 | 68.3824 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 57.622 | 54.878 | struthio_camelus_australis | ENSSCUP00000007240 | 49.7368 | 47.3684 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 70.7317 | 69.8171 | anser_brachyrhynchus | ENSABRP00000020543 | 84.6715 | 83.5766 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 29.878 | 29.5732 | gasterosteus_aculeatus | ENSGACP00000026825 | 94.2308 | 93.2692 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 53.0488 | 50 | gasterosteus_aculeatus | ENSGACP00000026824 | 82.4645 | 77.7251 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 43.5976 | 40.8537 | cynoglossus_semilaevis | ENSCSEP00000021219 | 80.791 | 75.7062 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 52.439 | 46.6463 | cynoglossus_semilaevis | ENSCSEP00000021231 | 76.7857 | 68.3036 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 83.2317 | 79.8781 | cynoglossus_semilaevis | ENSCSEP00000005055 | 85.5799 | 82.1317 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 29.5732 | 29.2683 | poecilia_formosa | ENSPFOP00000019454 | 83.6207 | 82.7586 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 53.6585 | 51.5244 | poecilia_formosa | ENSPFOP00000019456 | 56.7742 | 54.5161 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 90.2439 | myripristis_murdjan | ENSMMDP00005052606 | 73.8272 | 73.0864 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 86.5854 | 84.1463 | myripristis_murdjan | ENSMMDP00005025837 | 74.5407 | 72.4409 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 31.4024 | 29.878 | sander_lucioperca | ENSSLUP00000024800 | 85.8333 | 81.6667 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 50.6098 | 48.7805 | sander_lucioperca | ENSSLUP00000042214 | 74.4395 | 71.7489 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.6341 | sander_lucioperca | ENSSLUP00000049029 | 74.2647 | 72.0588 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 83.2317 | 80.4878 | strigops_habroptila | ENSSHBP00005020884 | 84.7826 | 81.9876 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 40.5488 | 38.1098 | amphiprion_ocellaris | ENSAOCP00000010593 | 68.2051 | 64.1026 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 18.2927 | 15.5488 | amphiprion_ocellaris | ENSAOCP00000007798 | 46.1538 | 39.2308 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 73.4756 | 73.4756 | terrapene_carolina_triunguis | ENSTMTP00000020985 | 74.1538 | 74.1538 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 42.9878 | 39.3293 | hucho_hucho | ENSHHUP00000089991 | 67.1429 | 61.4286 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.7683 | 90.5488 | hucho_hucho | ENSHHUP00000011172 | 71.4964 | 70.5463 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 28.0488 | 25.9146 | hucho_hucho | ENSHHUP00000071831 | 82.1429 | 75.8929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 51.2195 | 49.3902 | hucho_hucho | ENSHHUP00000053710 | 73.0435 | 70.4348 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 82.9268 | 80.7927 | hucho_hucho | ENSHHUP00000018140 | 77.937 | 75.9312 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 42.0732 | 39.3293 | hucho_hucho | ENSHHUP00000023343 | 76.2431 | 71.2707 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | betta_splendens | ENSBSLP00000039409 | 76.6585 | 74.6929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.4268 | 94.2073 | pygocentrus_nattereri | ENSPNAP00000029039 | 80.4627 | 79.4344 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 96.3415 | 95.4268 | pygocentrus_nattereri | ENSPNAP00000001619 | 81.2339 | 80.4627 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | anabas_testudineus | ENSATEP00000015111 | 76.6585 | 74.6929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 89.3293 | anabas_testudineus | ENSATEP00000004729 | 74.1936 | 72.7047 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 59.7561 | 50.9146 | ciona_savignyi | ENSCSAVP00000017661 | 63.2258 | 53.871 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 49.0854 | 48.1707 | seriola_lalandi_dorsalis | ENSSLDP00000004683 | 79.3103 | 77.8325 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.3781 | 89.6341 | seriola_lalandi_dorsalis | ENSSLDP00000021990 | 74.2647 | 72.0588 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 26.2195 | 23.4756 | seriola_lalandi_dorsalis | ENSSLDP00000027512 | 66.1538 | 59.2308 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 88.7195 | 85.6707 | labrus_bergylta | ENSLBEP00000009192 | 69.1211 | 66.7458 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 74.6951 | 69.8171 | labrus_bergylta | ENSLBEP00000034904 | 71.8475 | 67.1554 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | pundamilia_nyererei | ENSPNYP00000024379 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | mastacembelus_armatus | ENSMAMP00000015471 | 76.4706 | 74.5098 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.8537 | 89.0244 | mastacembelus_armatus | ENSMAMP00000046030 | 68.5057 | 67.1264 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | stegastes_partitus | ENSSPAP00000023628 | 76.6585 | 74.6929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 17.6829 | 15.8537 | oncorhynchus_tshawytscha | ENSOTSP00005058787 | 58 | 52 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 83.2317 | 81.4024 | oncorhynchus_tshawytscha | ENSOTSP00005028278 | 70.9091 | 69.3506 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 55.4878 | 50 | oncorhynchus_tshawytscha | ENSOTSP00005058784 | 71.6535 | 64.5669 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 18.5976 | 16.7683 | oncorhynchus_tshawytscha | ENSOTSP00005090668 | 54.4643 | 49.1071 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 48.4756 | 43.5976 | oncorhynchus_tshawytscha | ENSOTSP00005090617 | 72.9358 | 65.5963 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 90.2439 | oncorhynchus_tshawytscha | ENSOTSP00005016303 | 73.4644 | 72.7273 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 90.2439 | oncorhynchus_tshawytscha | ENSOTSP00005032609 | 73.4644 | 72.7273 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | acanthochromis_polyacanthus | ENSAPOP00000000457 | 76.6585 | 74.6929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 98.1707 | 97.2561 | aquila_chrysaetos_chrysaetos | ENSACCP00020012034 | 93.0636 | 92.1965 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.1585 | 88.1098 | nothobranchius_furzeri | ENSNFUP00015047527 | 72.7494 | 70.3163 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | cyprinodon_variegatus | ENSCVAP00000014372 | 80.4124 | 78.0928 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 94.8171 | 94.5122 | denticeps_clupeoides | ENSDCDP00000040566 | 77.5561 | 77.3067 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.9878 | 91.4634 | denticeps_clupeoides | ENSDCDP00000011325 | 75.1232 | 73.8916 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | larimichthys_crocea | ENSLCRP00005063801 | 76.6585 | 74.6929 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.8537 | 89.0244 | takifugu_rubripes | ENSTRUP00000067720 | 68.6636 | 67.2811 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 26.2195 | 22.8659 | ficedula_albicollis | ENSFALP00000017982 | 46.2366 | 40.3226 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 91.4634 | 87.8049 | hippocampus_comes | ENSHCOP00000021366 | 73.1707 | 70.2439 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 80.4878 | 75.3049 | hippocampus_comes | ENSHCOP00000013968 | 65.8354 | 61.596 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.7317 | 94.8171 | cyprinus_carpio_carpio | ENSCCRP00000151483 | 76.9608 | 76.2255 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.0732 | 91.4634 | cyprinus_carpio_carpio | ENSCCRP00000142251 | 73.8386 | 73.3496 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 92.6829 | 89.6341 | cyprinus_carpio_carpio | ENSCCRP00000115377 | 76.7677 | 74.2424 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.5 | 84.4512 | cyprinus_carpio_carpio | ENSCCRP00000152248 | 73.4015 | 70.844 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 52.439 | 51.2195 | coturnix_japonica | ENSCJPP00005023539 | 82.2967 | 80.3828 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 32.3171 | 27.439 | coturnix_japonica | ENSCJPP00005023540 | 61.6279 | 52.3256 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 44.8171 | 41.7683 | poecilia_latipinna | ENSPLAP00000029350 | 78.1915 | 72.8723 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 46.9512 | 44.8171 | poecilia_latipinna | ENSPLAP00000012171 | 84.6154 | 80.7692 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 74.0854 | 69.8171 | sinocyclocheilus_grahami | ENSSGRP00000047661 | 67.1271 | 63.2597 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.1951 | 85.3659 | sinocyclocheilus_grahami | ENSSGRP00000059819 | 74.0933 | 72.5389 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 46.0366 | 43.5976 | sinocyclocheilus_grahami | ENSSGRP00000055748 | 68.018 | 64.4144 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 16.4634 | 14.939 | sinocyclocheilus_grahami | ENSSGRP00000055749 | 42.1875 | 38.2812 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.8049 | 86.2805 | sinocyclocheilus_grahami | ENSSGRP00000040690 | 72.9114 | 71.6456 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 27.439 | 25.9146 | sinocyclocheilus_grahami | ENSSGRP00000055747 | 94.7368 | 89.4737 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.3781 | maylandia_zebra | ENSMZEP00005010177 | 76.6585 | 74.4472 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 47.8659 | 42.6829 | tetraodon_nigroviridis | ENSTNIP00000014788 | 74.0566 | 66.0377 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.939 | 88.1098 | paramormyrops_kingsleyae | ENSPKIP00000023477 | 76.2274 | 74.677 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 28.6585 | 26.5244 | paramormyrops_kingsleyae | ENSPKIP00000039954 | 83.1858 | 76.9911 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 54.2683 | 53.3537 | paramormyrops_kingsleyae | ENSPKIP00000028374 | 81.6514 | 80.2752 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 87.1951 | 85.6707 | scleropages_formosus | ENSSFOP00015018947 | 68.2578 | 67.0644 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 90.5488 | 87.8049 | scleropages_formosus | ENSSFOP00015050184 | 91.1043 | 88.3436 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 33.8415 | 31.0976 | leptobrachium_leishanense | ENSLLEP00000001291 | 89.5161 | 82.2581 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 72.2561 | 69.2073 | leptobrachium_leishanense | ENSLLEP00000001357 | 66.2011 | 63.4078 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 89.6341 | 86.5854 | scophthalmus_maximus | ENSSMAP00000030150 | 52.6882 | 50.8961 |
| ENSG00000153944 | homo_sapiens | ENSP00000284073 | 95.1219 | 92.6829 | oryzias_melastigma | ENSOMEP00000025152 | 80.4124 | 78.3505 |
Gene Ontology
| Go ID | Go term | No. evidence | Entries | Species | Category |
|---|---|---|---|---|---|
| GO:0003723 | enables RNA binding | 1 | HDA | Homo_sapiens(9606) | Function |
| GO:0003729 | enables mRNA binding | 1 | IBA | Homo_sapiens(9606) | Function |
| GO:0005515 | enables protein binding | 1 | IPI | Homo_sapiens(9606) | Function |
| GO:0005737 | is_active_in cytoplasm | 1 | IBA | Homo_sapiens(9606) | Component |
| GO:0005829 | located_in cytosol | 2 | IDA,TAS | Homo_sapiens(9606) | Component |
| GO:0005844 | part_of polysome | 1 | IBA | Homo_sapiens(9606) | Component |
| GO:0006417 | involved_in regulation of translation | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0007417 | involved_in central nervous system development | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0008266 | enables poly(U) RNA binding | 1 | IEA | Homo_sapiens(9606) | Function |
| GO:0042802 | enables identical protein binding | 1 | IPI | Homo_sapiens(9606) | Function |
| GO:0043231 | located_in intracellular membrane-bounded organelle | 1 | IDA | Homo_sapiens(9606) | Component |
| GO:0048864 | involved_in stem cell development | 1 | IEA | Homo_sapiens(9606) | Process |