Welcome to
RBP World!
Functions and Diseases
| RBP Type | Canonical_RBPs |
| Diseases | CancerCHIKVSARS-COV-OC43FlavivirusesHIVRVSARS-COV-2SINVDengueZika |
| Drug | N.A. |
| Main interacting RNAs | mRNA |
| Moonlighting functions | N.A. |
| Localizations | Stress granucle |
| BulkPerturb-seq | DataSet_01_172 DataSet_01_402 |
Description
| Ensembl ID | ENSG00000114867 | Gene ID | 1981 | Accession | 3296 |
| Symbol | EIF4G1 | Alias | P220;EIF4F;EIF4G;EIF4GI;PARK18;EIF-4G1 | Full Name | eukaryotic translation initiation factor 4 gamma 1 |
| Status | Confidence | Length | 20864 bases | Strand | Plus strand |
| Position | 3 : 184314495 - 184335358 | RNA binding domain | W2 , MIF4G , MA3 | ||
| Summary | The protein encoded by this gene is a component of the multi-subunit protein complex EIF4F. This complex facilitates the recruitment of mRNA to the ribosome, which is a rate-limiting step during the initiation phase of protein synthesis. The recognition of the mRNA cap and the ATP-dependent unwinding of 5'-terminal secondary structure is catalyzed by factors in this complex. The subunit encoded by this gene is a large scaffolding protein that contains binding sites for other members of the EIF4F complex. A domain at its N-terminus can also interact with the poly(A)-binding protein, which may mediate the circularization of mRNA during translation. Alternative splicing results in multiple transcript variants, some of which are derived from alternative promoter usage. [provided by RefSeq, Aug 2010] | ||||
RNA binding domains (RBDs)
| Protein | Domain | Pfam ID | E-value | Domain number |
|---|---|---|---|---|
| ENSP00000411826 | MA3 | PF02847 | 2.1e-29 | 1 |
| ENSP00000411826 | MIF4G | PF02854 | 1.7e-68 | 2 |
| ENSP00000411826 | W2 | PF02020 | 3.8e-18 | 3 |
| ENSP00000317600 | MA3 | PF02847 | 2.2e-29 | 1 |
| ENSP00000317600 | MIF4G | PF02854 | 1.8e-68 | 2 |
| ENSP00000317600 | W2 | PF02020 | 3.9e-18 | 3 |
| ENSP00000399858 | MA3 | PF02847 | 2.2e-29 | 1 |
| ENSP00000399858 | MIF4G | PF02854 | 1.8e-68 | 2 |
| ENSP00000399858 | W2 | PF02020 | 3.9e-18 | 3 |
| ENSP00000376320 | MA3 | PF02847 | 2.4e-29 | 1 |
| ENSP00000376320 | MIF4G | PF02854 | 1.9e-68 | 2 |
| ENSP00000376320 | W2 | PF02020 | 4.2e-18 | 3 |
| ENSP00000407682 | MA3 | PF02847 | 2.4e-29 | 1 |
| ENSP00000407682 | MIF4G | PF02854 | 1.9e-68 | 2 |
| ENSP00000407682 | W2 | PF02020 | 4.2e-18 | 3 |
| ENSP00000391935 | MA3 | PF02847 | 2.5e-29 | 1 |
| ENSP00000391935 | MIF4G | PF02854 | 2e-68 | 2 |
| ENSP00000391935 | W2 | PF02020 | 4.3e-18 | 3 |
| ENSP00000395974 | MA3 | PF02847 | 2.5e-29 | 1 |
| ENSP00000395974 | MIF4G | PF02854 | 2e-68 | 2 |
| ENSP00000395974 | W2 | PF02020 | 4.3e-18 | 3 |
| ENSP00000404754 | MA3 | PF02847 | 2.6e-29 | 1 |
| ENSP00000404754 | MIF4G | PF02854 | 2.1e-68 | 2 |
| ENSP00000404754 | W2 | PF02020 | 4.4e-18 | 3 |
| ENSP00000316879 | MA3 | PF02847 | 2.7e-29 | 1 |
| ENSP00000316879 | MIF4G | PF02854 | 2.1e-68 | 2 |
| ENSP00000316879 | W2 | PF02020 | 4.5e-18 | 3 |
| ENSP00000343450 | MA3 | PF02847 | 2.7e-29 | 1 |
| ENSP00000343450 | MIF4G | PF02854 | 2.1e-68 | 2 |
| ENSP00000343450 | W2 | PF02020 | 4.5e-18 | 3 |
| ENSP00000338020 | MA3 | PF02847 | 2.8e-29 | 1 |
| ENSP00000338020 | MIF4G | PF02854 | 2.1e-68 | 2 |
| ENSP00000338020 | W2 | PF02020 | 4.5e-18 | 3 |
| ENSP00000371767 | MA3 | PF02847 | 2.8e-29 | 1 |
| ENSP00000371767 | MIF4G | PF02854 | 2.1e-68 | 2 |
| ENSP00000371767 | W2 | PF02020 | 4.5e-18 | 3 |
| ENSP00000416255 | MA3 | PF02847 | 2.8e-29 | 1 |
| ENSP00000416255 | MIF4G | PF02854 | 2.1e-68 | 2 |
| ENSP00000416255 | W2 | PF02020 | 4.5e-18 | 3 |
| ENSP00000396888 | MA3 | PF02847 | 2e-09 | 1 |
| ENSP00000403269 | MIF4G | PF02854 | 2.6e-58 | 1 |
| ENSP00000398145 | MIF4G | PF02854 | 5.6e-42 | 1 |
| ENSP00000413159 | MIF4G | PF02854 | 8.4e-26 | 1 |
| ENSP00000391412 | MIF4G | PF02854 | 2.9e-18 | 1 |
| ENSP00000392908 | MIF4G | PF02854 | 7.6e-09 | 1 |
RNA binding proteomes (RBPomes)
| Pubmed ID | Full Name | Cell | Author | Time | Doi |
|---|
Literatures on RNA binding capacity
| Pubmed ID | Title | Author | Time | Journal |
|---|---|---|---|---|
| 24157614 | Depletion of hnRNP A2/B1 overrides the nuclear retention of the HIV-1 genomic RNA. | Heather Gordon | 2013-11-01 | RNA biology |
| 27668427 | Characterization of the Expression of the RNA Binding Protein eIF4G1 and Its Clinicopathological Correlation with Serous Ovarian Cancer. | Lanfang Li | 2016-01-01 | PloS one |
| 33510277 | A translation enhancer element from black beetle virus engages yeast eIF4G1 to drive cap-independent translation initiation. | M Brandon Trainor | 2021-01-28 | Scientific reports |
| 35291721 | Short and Sweet: Viral 5`-UTR as a Canonical and Non-Canonical Translation Initiation Switch. | M Brandon Trainor | 2021-01-01 | Journal of cellular immunology |
| 24183571 | Dynamic recognition of the mRNA cap by Saccharomyces cerevisiae eIF4E. | E Seán O'Leary | 2013-12-03 | Structure (London, England : 1993) |
| 28546148 | Translation initiation factor eIF4G1 preferentially binds yeast transcript leaders containing conserved oligo-uridine motifs. | Boris Zinshteyn | 2017-09-01 | RNA (New York, N.Y.) |
| 33396899 | Role of Host-Mediated Post-Translational Modifications (PTMs) in RNA Virus Pathogenesis. | Ramesh Kumar | 2020-12-30 | International journal of molecular sciences |
| 28738064 | Species A rotavirus NSP3 acquires its translation inhibitory function prior to stable dimer formation. | I Hugo Contreras-Treviño | 2017-01-01 | PloS one |
| 35018467 | Ribosomal leaky scanning through a translated uORF requires eIF4G2. | V Victoria Smirnova | 2022-01-25 | Nucleic acids research |
| 33523588 | Elevation of EIF4G1 promotes non-small cell lung cancer progression by activating mTOR signalling. | Ying Lu | 2021-03-01 | Journal of cellular and molecular medicine |
| 35041803 | Direct analysis of ribosome targeting illuminates thousand-fold regulation of translation initiation. | O Rachel Niederer | 2022-03-16 | Cell systems |
| 31157589 | RGG-motif self-association regulates eIF4G-binding translation repressor protein Scd6. | Gopalakrishna Poornima | 2019-09-01 | RNA biology |
| 29603493 | Long non-coding RNA-CTD-2108O9.1 represses breast cancer metastasis by influencing leukemia inhibitory factor receptor. | Mozhi Wang | 2018-06-01 | Cancer science |
Transcripts
| Name | Transcript ID | bp | Protein | Translation ID |
|---|---|---|---|---|
| EIF4G1-204 | ENST00000352767 | 5484 | 1606aa | ENSP00000338020 |
| EIF4G1-209 | ENST00000414031 | 5569 | 1559aa | ENSP00000391935 |
| EIF4G1-206 | ENST00000392537 | 5229 | 1512aa | ENSP00000376320 |
| EIF4G1-223 | ENST00000444134 | 558 | 127aa | ENSP00000407244 |
| EIF4G1-222 | ENST00000442406 | 5129 | 50aa | ENSP00000400351 |
| EIF4G1-226 | ENST00000450424 | 2658 | 833aa | ENSP00000391412 |
| EIF4G1-210 | ENST00000421110 | 2755 | 869aa | ENSP00000413159 |
| EIF4G1-219 | ENST00000435046 | 5042 | 1577aa | ENSP00000404754 |
| EIF4G1-205 | ENST00000382330 | 5129 | 1606aa | ENSP00000371767 |
| EIF4G1-208 | ENST00000413967 | 2808 | 71aa | ENSP00000390755 |
| EIF4G1-213 | ENST00000426123 | 2907 | 903aa | ENSP00000403269 |
| EIF4G1-202 | ENST00000346169 | 5405 | 1599aa | ENSP00000316879 |
| EIF4G1-203 | ENST00000350481 | 4990 | 1435aa | ENSP00000317600 |
| EIF4G1-227 | ENST00000455679 | 542 | 87aa | ENSP00000415842 |
| EIF4G1-237 | ENST00000485712 | 474 | No protein | - |
| EIF4G1-220 | ENST00000440448 | 578 | 143aa | ENSP00000407240 |
| EIF4G1-214 | ENST00000427141 | 730 | 92aa | ENSP00000411214 |
| EIF4G1-216 | ENST00000427845 | 5038 | 1513aa | ENSP00000407682 |
| EIF4G1-201 | ENST00000342981 | 5667 | 1600aa | ENSP00000343450 |
| EIF4G1-212 | ENST00000424196 | 5653 | 1606aa | ENSP00000416255 |
| EIF4G1-228 | ENST00000456033 | 694 | 160aa | ENSP00000415943 |
| EIF4G1-207 | ENST00000411531 | 4683 | 1560aa | ENSP00000395974 |
| EIF4G1-224 | ENST00000444861 | 2384 | 757aa | ENSP00000398145 |
| EIF4G1-236 | ENST00000484862 | 590 | No protein | - |
| EIF4G1-239 | ENST00000676206 | 702 | 103aa | ENSP00000502064 |
| EIF4G1-221 | ENST00000441154 | 4311 | 1436aa | ENSP00000399858 |
| EIF4G1-217 | ENST00000428387 | 524 | 143aa | ENSP00000411707 |
| EIF4G1-218 | ENST00000434061 | 5026 | 1404aa | ENSP00000411826 |
| EIF4G1-215 | ENST00000427607 | 596 | 163aa | ENSP00000409545 |
| EIF4G1-229 | ENST00000457456 | 983 | 285aa | ENSP00000399969 |
| EIF4G1-238 | ENST00000493299 | 559 | No protein | - |
| EIF4G1-240 | ENST00000676453 | 4590 | 525aa | ENSP00000501695 |
| EIF4G1-225 | ENST00000448284 | 677 | 226aa | ENSP00000392908 |
| EIF4G1-232 | ENST00000466311 | 573 | No protein | - |
| EIF4G1-235 | ENST00000482303 | 493 | No protein | - |
| EIF4G1-211 | ENST00000422614 | 750 | 180aa | ENSP00000396888 |
| EIF4G1-230 | ENST00000460829 | 1804 | No protein | - |
| EIF4G1-233 | ENST00000475721 | 1271 | No protein | - |
| EIF4G1-231 | ENST00000464548 | 637 | No protein | - |
| EIF4G1-234 | ENST00000478291 | 564 | No protein | - |
Phenotypes
| ensgID | Trait | pValue | Pubmed ID |
|---|---|---|---|
| 77582 | Menarche | 3E-12 | 25231870 |
GWAS
| ensgID | SNP | Chromosome | Position | Trait | PubmedID | Or or BEAT | EFO ID |
|---|
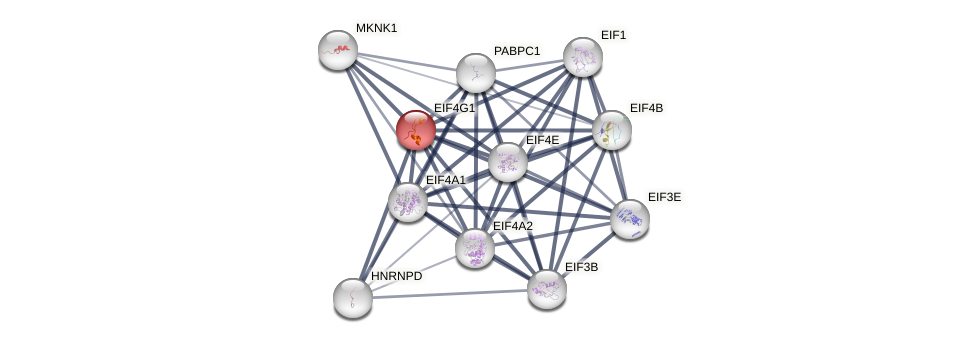
Protein-Protein Interaction (PPI)
Paralogs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| ENSG00000114867 | homo_sapiens | ENSP00000473510 | 57.0993 | 46.1304 | homo_sapiens | ENSP00000316879 | 58.5991 | 47.3421 |
| ENSG00000114867 | homo_sapiens | ENSP00000340281 | 44.8732 | 28.8864 | homo_sapiens | ENSP00000316879 | 25.4534 | 16.3852 |
Orthologs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.7473 | 95.3096 | nomascus_leucogenys | ENSNLEP00000035819 | 98.267 | 97.8177 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 99.4371 | 99.3121 | pan_paniscus | ENSPPAP00000040280 | 99.0037 | 98.8792 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 99.187 | 99.1245 | pongo_abelii | ENSPPYP00000023530 | 98.7547 | 98.6924 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 97.2483 | 97.1232 | pan_troglodytes | ENSPTRP00000026995 | 99.7434 | 99.6151 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 99.3121 | 99.187 | gorilla_gorilla | ENSGGOP00000013587 | 99.0643 | 98.9395 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 84.6779 | 79.2996 | ornithorhynchus_anatinus | ENSOANP00000051357 | 83.5287 | 78.2233 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 88.3677 | 83.4897 | sarcophilus_harrisii | ENSSHAP00000031983 | 87.33 | 82.5093 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 83.1144 | 78.8618 | notamacropus_eugenii | ENSMEUP00000009907 | 83.3752 | 79.1092 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.6104 | 74.7342 | choloepus_hoffmanni | ENSCHOP00000001523 | 83.3333 | 81.2925 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 88.4928 | 86.8668 | dasypus_novemcinctus | ENSDNOP00000024671 | 95.4147 | 93.6615 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 68.9181 | 67.4171 | erinaceus_europaeus | ENSEEUP00000003599 | 84.5092 | 82.6687 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 65.3533 | 63.0394 | echinops_telfairi | ENSETEP00000003467 | 67.4194 | 65.0323 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 97.9987 | 97.3108 | callithrix_jacchus | ENSCJAP00000027887 | 97.6324 | 96.947 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.8118 | 98.6241 | cercocebus_atys | ENSCATP00000002195 | 98.5037 | 98.3167 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.6241 | 98.4365 | macaca_fascicularis | ENSMFAP00000035962 | 98.1943 | 98.0075 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.6867 | 98.4991 | macaca_mulatta | ENSMMUP00000014338 | 98.3791 | 98.192 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.6867 | 98.4991 | macaca_nemestrina | ENSMNEP00000028468 | 98.3791 | 98.192 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.9368 | 98.5616 | papio_anubis | ENSPANP00000028132 | 98.567 | 98.1931 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.9994 | 98.7492 | mandrillus_leucophaeus | ENSMLEP00000022765 | 99.0613 | 98.811 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.0488 | 76.3602 | tursiops_truncatus | ENSTTRP00000010666 | 81.1971 | 79.4405 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.9356 | 95.0594 | vulpes_vulpes | ENSVVUP00000016524 | 96.2733 | 94.4099 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.9337 | 91.0569 | mus_spretus | MGP_SPRETEiJ_P0042837 | 93.875 | 91 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.4978 | 95.3721 | equus_asinus | ENSEASP00005020377 | 95.7196 | 94.603 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.6223 | 93.8086 | capra_hircus | ENSCHIP00000005945 | 95.1462 | 93.3416 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 90.8068 | 88.4303 | loxodonta_africana | ENSLAFP00000004595 | 94.1634 | 91.6991 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.748 | 94.9343 | panthera_leo | ENSPLOP00000013558 | 96.087 | 94.2857 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.873 | 94.7467 | ailuropoda_melanoleuca | ENSAMEP00000018112 | 96.2112 | 94.0994 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.8099 | 94.1839 | bos_indicus_hybrid | ENSBIXP00005039341 | 95.3329 | 93.715 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.6229 | 94.9343 | oryctolagus_cuniculus | ENSOCUP00000032992 | 96.0821 | 94.403 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.6854 | 95.4972 | equus_caballus | ENSECAP00000006848 | 96.324 | 95.1402 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.873 | 94.9969 | canis_lupus_familiaris | ENSCAFP00845038624 | 96.2112 | 94.3478 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.748 | 94.9343 | panthera_pardus | ENSPPRP00000014602 | 96.087 | 94.2857 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.1851 | 94.3715 | marmota_marmota_marmota | ENSMMMP00000008506 | 95.4094 | 93.6104 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.4353 | 94.6842 | balaenoptera_musculus | ENSBMSP00010010456 | 95.9552 | 94.2128 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.1207 | 90.4941 | cavia_porcellus | ENSCPOP00000006613 | 94.3002 | 91.6403 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.8105 | 94.9343 | felis_catus | ENSFCAP00000007218 | 96.5087 | 94.6384 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 82.4265 | 80.2376 | ursus_americanus | ENSUAMP00000003923 | 95.1625 | 92.6354 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 59.3496 | 49.656 | monodelphis_domestica | ENSMODP00000032477 | 63.0565 | 52.7575 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 77.8612 | 72.4828 | monodelphis_domestica | ENSMODP00000046948 | 82.3413 | 76.6534 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.4346 | 93.8086 | bison_bison_bison | ENSBBBP00000021051 | 94.9595 | 93.3416 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.3102 | 94.8093 | monodon_monoceros | ENSMMNP00015005326 | 95.8307 | 94.3373 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 92.8705 | 91.4321 | sus_scrofa | ENSSSCP00000052793 | 92.0645 | 90.6386 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.4353 | 95.3721 | microcebus_murinus | ENSMICP00000051313 | 96.0748 | 95.0156 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 11.8824 | 11.8824 | octodon_degus | ENSODEP00000017262 | 94.0594 | 94.0594 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.5478 | 73.8587 | octodon_degus | ENSODEP00000018820 | 93.0798 | 89.8099 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 62.8518 | 61.0381 | jaculus_jaculus | ENSJJAP00000007881 | 93.8375 | 91.1298 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 12.07 | 12.0075 | jaculus_jaculus | ENSJJAP00000019280 | 92.3445 | 91.866 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.7473 | 93.8712 | ovis_aries_rambouillet | ENSOARP00020003903 | 95.2707 | 93.4039 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.6854 | 94.6216 | ursus_maritimus | ENSUMAP00000006899 | 96.4442 | 94.3855 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 90.4315 | 88.4928 | ochotona_princeps | ENSOPRP00000013632 | 89.8695 | 87.9428 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.3102 | 94.8093 | delphinapterus_leucas | ENSDLEP00000013051 | 95.8307 | 94.3373 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 92.4953 | 90.8068 | physeter_catodon | ENSPCTP00005025374 | 95.7282 | 93.9806 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.1213 | 91.2445 | mus_caroli | MGP_CAROLIEiJ_P0040980 | 94.0625 | 91.1875 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.9337 | 91.2445 | mus_musculus | ENSMUSP00000111120 | 93.875 | 91.1875 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 12.1951 | 12.1326 | dipodomys_ordii | ENSDORP00000019242 | 93.3014 | 92.823 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.1113 | 76.6729 | dipodomys_ordii | ENSDORP00000009612 | 95.6355 | 93.8744 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 91.182 | 87.9925 | mustela_putorius_furo | ENSMPUP00000002800 | 85.967 | 82.9599 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 97.4984 | 96.748 | aotus_nancymaae | ENSANAP00000024217 | 96.5325 | 95.7895 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.1238 | 97.3734 | saimiri_boliviensis_boliviensis | ENSSBOP00000019560 | 97.6961 | 96.9489 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 40.838 | 40.025 | vicugna_pacos | ENSVPAP00000008378 | 46.9446 | 46.0101 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.1226 | 95.935 | chlorocebus_sabaeus | ENSCSAP00000009860 | 98.2109 | 98.0192 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 62.2889 | 60.6629 | tupaia_belangeri | ENSTBEP00000001726 | 73.7232 | 71.7987 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.98 | 66.2289 | mesocricetus_auratus | ENSMAUP00000024673 | 94.604 | 92.1671 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 12.0075 | 11.8199 | mesocricetus_auratus | ENSMAUP00000006919 | 91.866 | 90.4306 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.1839 | 92.5578 | phocoena_sinus | ENSPSNP00000004700 | 80.2772 | 78.8913 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.3727 | 94.8093 | camelus_dromedarius | ENSCDRP00005004501 | 95.9527 | 94.396 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.8105 | 95.1219 | carlito_syrichta | ENSTSYP00000031632 | 96.2687 | 94.5896 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.9969 | 93.0582 | panthera_tigris_altaica | ENSPTIP00000020366 | 94.4652 | 92.5373 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.6836 | 90.8693 | rattus_norvegicus | ENSRNOP00000067492 | 91.2858 | 88.5436 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.3727 | 95.3096 | prolemur_simus | ENSPSMP00000030071 | 95.9527 | 94.8941 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.0588 | 91.995 | microtus_ochrogaster | ENSMOCP00000017575 | 94.0588 | 91.995 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 8.88055 | 8.56785 | microtus_ochrogaster | ENSMOCP00000000658 | 83.0409 | 80.117 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.0413 | 62.7892 | sorex_araneus | ENSSARP00000003008 | 69.9801 | 66.5341 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.8086 | 91.6198 | peromyscus_maniculatus_bairdii | ENSPEMP00000018801 | 93.633 | 91.4482 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.9975 | 94.3715 | bos_mutus | ENSBMUP00000026121 | 95.5196 | 93.9017 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 5.81613 | 5.81613 | mus_spicilegus | ENSMSIP00000018420 | 93.9394 | 93.9394 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.9962 | 91.9325 | cricetulus_griseus_chok1gshd | ENSCGRP00001026951 | 94.1729 | 92.1053 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.434 | 92.3702 | pteropus_vampyrus | ENSPVAP00000004591 | 93.7888 | 91.7391 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 91.3071 | 88.9931 | myotis_lucifugus | ENSMLUP00000011674 | 93.7099 | 91.335 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 88.5553 | 83.4897 | vombatus_ursinus | ENSVURP00010030320 | 87.6238 | 82.6114 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 92.3077 | 89.9312 | rhinolophus_ferrumequinum | ENSRFEP00010029649 | 92.7718 | 90.3834 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.4353 | 94.434 | neovison_vison | ENSNVIP00000019185 | 95.7169 | 93.7306 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.8724 | 94.2464 | bos_taurus | ENSBTAP00000032150 | 95.3951 | 93.7772 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.7467 | 92.1826 | heterocephalus_glaber_female | ENSHGLP00000019408 | 93.9244 | 91.3825 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 35.0844 | 33.3959 | heterocephalus_glaber_female | ENSHGLP00000012183 | 90.9238 | 86.5478 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 97.6861 | 97.1857 | rhinopithecus_bieti | ENSRBIP00000026838 | 98.5489 | 98.0442 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.9356 | 95.0594 | canis_lupus_dingo | ENSCAFP00020002993 | 96.6937 | 94.8222 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.5603 | 95.3096 | sciurus_vulgaris | ENSSVLP00005016766 | 95.7816 | 94.5409 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 88.6179 | 83.3646 | phascolarctos_cinereus | ENSPCIP00000034667 | 87.9032 | 82.6923 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 86.8043 | 83.3021 | procavia_capensis | ENSPCAP00000013803 | 89.2605 | 85.6592 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 10.3189 | 10.3189 | nannospalax_galili | ENSNGAP00000023689 | 97.0588 | 97.0588 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.6729 | 73.546 | nannospalax_galili | ENSNGAP00000010844 | 89.4238 | 85.7768 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.9856 | 74.7967 | nannospalax_galili | ENSNGAP00000021347 | 94.4018 | 91.7178 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 93.6836 | 92.1201 | bos_grunniens | ENSBGRP00000008044 | 95.5967 | 94.0013 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 94.5591 | 91.5572 | chinchilla_lanigera | ENSCLAP00000018902 | 93.913 | 90.9317 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.6848 | 93.621 | cervus_hanglu_yarkandensis | ENSCHYP00000012047 | 95.2085 | 93.1549 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.4346 | 94.0588 | catagonus_wagneri | ENSCWAP00000019493 | 94.7826 | 93.4161 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.2477 | 94.5591 | ictidomys_tridecemlineatus | ENSSTOP00000026974 | 95.5307 | 93.8548 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.4978 | 95.3096 | otolemur_garnettii | ENSOGAP00000013701 | 94.9538 | 93.7846 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.2489 | 97.3734 | cebus_imitator | ENSCCAP00000038938 | 97.8816 | 97.0093 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.3102 | 94.6216 | urocitellus_parryii | ENSUPAP00010003925 | 95.5928 | 93.9168 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 95.8099 | 93.9962 | moschus_moschiferus | ENSMMSP00000015681 | 95.3329 | 93.5283 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 98.8743 | 98.4365 | rhinopithecus_roxellana | ENSRROP00000001426 | 98.5661 | 98.1297 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 4.75297 | 4.69043 | mus_pahari | MGP_PahariEiJ_P0028352 | 96.2025 | 94.9367 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 96.6854 | 95.6223 | propithecus_coquereli | ENSPCOP00000028244 | 96.324 | 95.2648 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 21.7636 | 15.3846 | ciona_intestinalis | ENSCINP00000010668 | 50.508 | 35.7039 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 37.4609 | 29.9562 | eptatretus_burgeri | ENSEBUP00000018528 | 48.1899 | 38.5358 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 63.3521 | 52.6579 | callorhinchus_milii | ENSCMIP00000020027 | 68.3075 | 56.7768 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 77.6735 | 66.6041 | latimeria_chalumnae | ENSLACP00000005454 | 75.7317 | 64.939 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.3559 | 59.7248 | lepisosteus_oculatus | ENSLOCP00000010629 | 64.8917 | 55.8806 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.4816 | 59.6623 | clupea_harengus | ENSCHAP00000019943 | 62.6808 | 53.059 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 20.0125 | 16.4478 | danio_rerio | ENSDARP00000152393 | 74.7664 | 61.4486 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 23.3896 | 19.0744 | carassius_auratus | ENSCARP00000097407 | 72.3404 | 58.9942 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.419 | 59.3496 | astyanax_mexicanus | ENSAMXP00000019918 | 66.0023 | 55.6272 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.5422 | 57.536 | ictalurus_punctatus | ENSIPUP00000021766 | 68.268 | 58.1542 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 63.0394 | 52.0951 | ictalurus_punctatus | ENSIPUP00000022834 | 63.3962 | 52.3899 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.5435 | 57.7236 | esox_lucius | ENSELUP00000004979 | 62.5422 | 51.9123 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 65.2283 | 54.9093 | electrophorus_electricus | ENSEEEP00000030746 | 64.4623 | 54.2645 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 23.202 | 14.384 | oncorhynchus_kisutch | ENSOKIP00005083623 | 48.3073 | 29.9479 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 23.3271 | 14.384 | oncorhynchus_kisutch | ENSOKIP00005011298 | 47.8819 | 29.525 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 23.3896 | 14.4465 | oncorhynchus_mykiss | ENSOMYP00000020097 | 47.8261 | 29.5396 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 21.2633 | 13.571 | oncorhynchus_mykiss | ENSOMYP00000115529 | 47.5524 | 30.3496 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 14.5716 | 9.00563 | salmo_salar | ENSSSAP00000132071 | 45.5969 | 28.18 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 26.5791 | 16.4478 | salmo_salar | ENSSSAP00000096569 | 46.2459 | 28.6181 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 26.5166 | 16.3852 | salmo_trutta | ENSSTUP00000051767 | 47.216 | 29.1759 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 22.9518 | 14.0088 | salmo_trutta | ENSSTUP00000055551 | 47.8488 | 29.2047 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 68.1676 | 55.222 | fundulus_heteroclitus | ENSFHEP00000020984 | 64.497 | 52.2485 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.7292 | 54.2839 | poecilia_reticulata | ENSPREP00000004383 | 62.2157 | 50.6122 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.9168 | 54.3465 | xiphophorus_maculatus | ENSXMAP00000040642 | 62.6464 | 50.8782 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.354 | 55.4096 | oryzias_latipes | ENSORLP00000035606 | 61.8659 | 51.6618 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.9187 | 56.7229 | cyclopterus_lumpus | ENSCLMP00005032210 | 60.043 | 48.7111 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 65.2908 | 54.5966 | oreochromis_niloticus | ENSONIP00000061045 | 62.069 | 51.9025 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.479 | 55.4096 | haplochromis_burtoni | ENSHBUP00000023006 | 64.0747 | 53.4057 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.354 | 55.3471 | astatotilapia_calliptera | ENSACLP00000035122 | 61.8298 | 51.5734 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.8562 | 58.4115 | sparus_aurata | ENSSAUP00010017129 | 62.3674 | 52.1496 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.8568 | 59.7248 | lates_calcarifer | ENSLCAP00010028278 | 66.6863 | 56.2095 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 73.9837 | 63.9149 | xenopus_tropicalis | ENSXETP00000098195 | 73.5239 | 63.5177 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 79.1119 | 70.8568 | chrysemys_picta_bellii | ENSCPBP00000021647 | 77.9901 | 69.852 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.2933 | 61.3508 | sphenodon_punctatus | ENSSPUP00000017981 | 74.8649 | 66.2838 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.6729 | 67.6673 | crocodylus_porosus | ENSCPRP00005016483 | 76.1964 | 67.2467 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.9174 | 59.4747 | laticauda_laticaudata | ENSLLTP00000018002 | 74.4856 | 65.2263 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 72.4203 | 63.0394 | notechis_scutatus | ENSNSUP00000011588 | 76.0341 | 66.1852 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 74.6717 | 65.6035 | pseudonaja_textilis | ENSPTXP00000023964 | 76.834 | 67.5032 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 60.1626 | 52.6579 | anas_platyrhynchos_platyrhynchos | ENSAPLP00000016638 | 68.3239 | 59.8011 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.2364 | 69.3559 | gallus_gallus | ENSGALP00010024889 | 75.9563 | 67.3345 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.3602 | 68.0425 | meleagris_gallopavo | ENSMGAP00000033711 | 78.876 | 70.2842 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.6742 | 69.5435 | serinus_canaria | ENSSCAP00000019568 | 77.943 | 68.8971 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.1726 | 66.7917 | parus_major | ENSPMJP00000001969 | 74.1327 | 65.003 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 77.9237 | 69.2308 | anolis_carolinensis | ENSACAP00000002190 | 76.9611 | 68.3755 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 65.9162 | 54.7217 | neolamprologus_brichardi | ENSNBRP00000019842 | 66.2893 | 55.0314 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 73.0457 | 64.2276 | naja_naja | ENSNNAP00000028402 | 75.8934 | 66.7316 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 75.3596 | 67.5422 | podarcis_muralis | ENSPMRP00000009965 | 78.5016 | 70.3583 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 75.8599 | 68.0425 | geospiza_fortis | ENSGFOP00000020355 | 78.4605 | 70.3752 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.7992 | 69.2933 | taeniopygia_guttata | ENSTGUP00000019436 | 76.7824 | 67.5198 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 72.7955 | 62.7267 | erpetoichthys_calabaricus | ENSECRP00000007670 | 71.719 | 61.7991 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 68.2301 | 56.4103 | kryptolebias_marmoratus | ENSKMAP00000015773 | 63.9133 | 52.8412 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 77.611 | 69.4184 | salvator_merianae | ENSSMRP00000030741 | 77.3209 | 69.1589 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 62.9769 | 52.7205 | oryzias_javanicus | ENSOJAP00000040634 | 66.0328 | 55.2787 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.167 | 56.5353 | amphilophus_citrinellus | ENSACIP00000019038 | 65.8896 | 55.4601 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.7311 | 58.4115 | dicentrarchus_labrax | ENSDLAP00005072268 | 62.5701 | 52.413 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.0419 | 56.5979 | amphiprion_percula | ENSAPEP00000010140 | 69.7463 | 58.8809 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 53.2208 | 44.9031 | oryzias_sinensis | ENSOSIP00000047757 | 61.6667 | 52.029 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 18.7617 | 15.9475 | pelodiscus_sinensis | ENSPSIP00000018702 | 87.9765 | 74.7801 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 20.5754 | 18.0738 | pelodiscus_sinensis | ENSPSIP00000018678 | 63.2692 | 55.5769 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 79.237 | 70.7317 | gopherus_evgoodei | ENSGEVP00005006478 | 78.7935 | 70.3358 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.9869 | 70.6066 | chelonoidis_abingdonii | ENSCABP00000017418 | 78.7898 | 70.4304 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 71.5447 | 59.975 | seriola_dumerili | ENSSDUP00000012076 | 66.3188 | 55.5942 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.6685 | 58.0363 | cottoperca_gobio | ENSCGOP00000018020 | 63.9862 | 53.3027 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 74.1088 | 65.3533 | struthio_camelus_australis | ENSSCUP00000019557 | 76.998 | 67.9012 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.0488 | 69.2933 | anser_brachyrhynchus | ENSABRP00000001537 | 76.0049 | 67.4787 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.2308 | 57.4734 | gasterosteus_aculeatus | ENSGACP00000017882 | 69.058 | 57.33 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 64.5403 | 53.2833 | cynoglossus_semilaevis | ENSCSEP00000026268 | 70.3956 | 58.1173 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 65.7286 | 53.7211 | poecilia_formosa | ENSPFOP00000009203 | 65.2795 | 53.354 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.7317 | 60.1001 | myripristis_murdjan | ENSMMDP00005031961 | 67.7245 | 57.5449 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.7924 | 57.0982 | sander_lucioperca | ENSSLUP00000052426 | 63.8022 | 53.7375 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 76.4853 | 67.6048 | strigops_habroptila | ENSSHBP00005018211 | 76.9666 | 68.0302 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.9174 | 57.5985 | amphiprion_ocellaris | ENSAOCP00000006904 | 68.6473 | 58.2174 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 79.237 | 70.8568 | terrapene_carolina_triunguis | ENSTMTP00000015055 | 78.7935 | 70.4602 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 26.7667 | 16.3227 | hucho_hucho | ENSHHUP00000034894 | 47.4501 | 28.9357 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 5.62852 | 2.75172 | hucho_hucho | ENSHHUP00000076553 | 38.6266 | 18.8841 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 68.6679 | 57.0982 | betta_splendens | ENSBSLP00000043381 | 63.9115 | 53.1432 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 31.207 | 26.5791 | pygocentrus_nattereri | ENSPNAP00000031065 | 78.2132 | 66.6144 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.3565 | 58.7867 | anabas_testudineus | ENSATEP00000016135 | 66.9244 | 55.9191 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 71.4822 | 59.8499 | seriola_lalandi_dorsalis | ENSSLDP00000008831 | 64.9801 | 54.4059 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 29.6435 | 24.828 | labrus_bergylta | ENSLBEP00000024240 | 77.8325 | 65.1888 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.5416 | 55.4722 | pundamilia_nyererei | ENSPNYP00000015012 | 63.7507 | 53.1456 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 68.9181 | 56.9731 | mastacembelus_armatus | ENSMAMP00000018624 | 65.8697 | 54.4531 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 27.5172 | 23.6398 | stegastes_partitus | ENSSPAP00000028506 | 80.8824 | 69.4853 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 15.0094 | 9.13071 | oncorhynchus_tshawytscha | ENSOTSP00005005955 | 43.956 | 26.7399 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 12.5704 | 7.75485 | oncorhynchus_tshawytscha | ENSOTSP00005047901 | 43.0407 | 26.5525 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 23.2645 | 14.3215 | oncorhynchus_tshawytscha | ENSOTSP00005077549 | 48.4375 | 29.8177 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 17.8862 | 14.4465 | oncorhynchus_tshawytscha | ENSOTSP00005093296 | 80.791 | 65.2542 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.3559 | 58.8493 | acanthochromis_polyacanthus | ENSAPOP00000033054 | 66.13 | 56.1121 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 73.9837 | 65.6035 | aquila_chrysaetos_chrysaetos | ENSACCP00020004836 | 73.8452 | 65.4807 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.4165 | 54.7842 | nothobranchius_furzeri | ENSNFUP00015026080 | 60.1359 | 49.6036 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.9794 | 54.0963 | cyprinodon_variegatus | ENSCVAP00000021910 | 63.2231 | 51.0626 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.9812 | 59.2245 | denticeps_clupeoides | ENSDCDP00000025372 | 67.5725 | 57.186 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 22.7642 | 19.6373 | denticeps_clupeoides | ENSDCDP00000027523 | 81.6143 | 70.4036 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 71.1695 | 59.0369 | larimichthys_crocea | ENSLCRP00005061721 | 66.5108 | 55.1724 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.606 | 55.9725 | takifugu_rubripes | ENSTRUP00000076473 | 64.2239 | 51.6445 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.1739 | 69.0432 | ficedula_albicollis | ENSFALP00000008430 | 78.125 | 69 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.9794 | 55.3471 | hippocampus_comes | ENSHCOP00000010860 | 64.2857 | 53.1213 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 67.0419 | 56.4103 | cyprinus_carpio_carpio | ENSCCRP00000162592 | 64.1532 | 53.9797 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.9794 | 56.9106 | cyprinus_carpio_carpio | ENSCCRP00000011695 | 63.9403 | 54.3284 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 78.4866 | 69.4809 | coturnix_japonica | ENSCJPP00005016768 | 77.0411 | 68.2013 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.7917 | 54.5966 | poecilia_latipinna | ENSPLAP00000031593 | 63.4581 | 51.8717 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 29.3308 | 24.7655 | poecilia_latipinna | ENSPLAP00000017949 | 68.8693 | 58.1498 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.9794 | 56.4728 | sinocyclocheilus_grahami | ENSSGRP00000080241 | 66.729 | 56.2617 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 63.3521 | 52.783 | sinocyclocheilus_grahami | ENSSGRP00000022364 | 62.5695 | 52.1309 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.354 | 55.3471 | maylandia_zebra | ENSMZEP00005034034 | 63.8772 | 53.2812 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 12.8205 | 9.31832 | tetraodon_nigroviridis | ENSTNIP00000011083 | 67.8808 | 49.3377 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.6692 | 60.1626 | paramormyrops_kingsleyae | ENSPKIP00000034027 | 68.3606 | 58.1972 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 70.8568 | 60.2251 | scleropages_formosus | ENSSFOP00015029972 | 68.4592 | 58.1873 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 71.6073 | 62.2889 | leptobrachium_leishanense | ENSLLEP00000025309 | 70.1163 | 60.992 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 69.6685 | 58.4115 | scophthalmus_maximus | ENSSMAP00000000314 | 62.9734 | 52.7982 |
| ENSG00000114867 | homo_sapiens | ENSP00000316879 | 66.7292 | 55.9099 | oryzias_melastigma | ENSOMEP00000036208 | 64.0456 | 53.6615 |
Gene Ontology
| Go ID | Go term | No. evidence | Entries | Species | Category |
|---|---|---|---|---|---|
| GO:0001662 | involved_in behavioral fear response | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0002191 | involved_in cap-dependent translational initiation | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0003723 | enables RNA binding | 2 | HDA,TAS | Homo_sapiens(9606) | Function |
| GO:0003729 | enables mRNA binding | 2 | IBA,IDA | Homo_sapiens(9606) | Function |
| GO:0003743 | enables translation initiation factor activity | 3 | IBA,IDA,TAS | Homo_sapiens(9606) | Function |
| GO:0005515 | enables protein binding | 1 | IPI | Homo_sapiens(9606) | Function |
| GO:0005524 | enables ATP binding | 1 | IDA | Homo_sapiens(9606) | Function |
| GO:0005634 | located_in nucleus | 2 | IDA,IMP | Homo_sapiens(9606) | Component |
| GO:0005737 | located_in cytoplasm | 2 | IDA,IMP | Homo_sapiens(9606) | Component |
| GO:0005829 | located_in cytosol | 2 | IDA,TAS | Homo_sapiens(9606) | Component |
| GO:0005844 | part_of polysome | 1 | IMP | Homo_sapiens(9606) | Component |
| GO:0006412 | involved_in translation | 2 | IMP,IPI | Homo_sapiens(9606) | Process |
| GO:0006413 | involved_in translational initiation | 3 | IDA,NAS,TAS | Homo_sapiens(9606) | Process |
| GO:0006446 | involved_in regulation of translational initiation | 2 | IMP,NAS | Homo_sapiens(9606) | Process |
| GO:0008135 | enables translation factor activity, RNA binding | 1 | IMP | Homo_sapiens(9606) | Function |
| GO:0008190 | enables eukaryotic initiation factor 4E binding | 2 | IDA,TAS | Homo_sapiens(9606) | Function |
| GO:0009059 | involved_in macromolecule biosynthetic process | 1 | IGI | Homo_sapiens(9606) | Process |
| GO:0010494 | located_in cytoplasmic stress granule | 1 | IDA | Homo_sapiens(9606) | Component |
| GO:0010507 | involved_in negative regulation of autophagy | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0010801 | involved_in negative regulation of peptidyl-threonine phosphorylation | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0016020 | located_in membrane | 1 | HDA | Homo_sapiens(9606) | Component |
| GO:0016281 | part_of eukaryotic translation initiation factor 4F complex | 3 | IBA,IDA,NAS | Homo_sapiens(9606) | Component |
| GO:0030182 | involved_in neuron differentiation | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0030307 | involved_in positive regulation of cell growth | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0030324 | involved_in lung development | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0031369 | enables translation initiation factor binding | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0031669 | involved_in cellular response to nutrient levels | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0033138 | involved_in positive regulation of peptidyl-serine phosphorylation | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0035278 | involved_in miRNA-mediated gene silencing by inhibition of translation | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:0036493 | involved_in positive regulation of translation in response to endoplasmic reticulum stress | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0042802 | enables identical protein binding | 1 | IPI | Homo_sapiens(9606) | Function |
| GO:0045471 | involved_in response to ethanol | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0045666 | involved_in positive regulation of neuron differentiation | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0051247 | involved_in positive regulation of protein metabolic process | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0060090 | enables molecular adaptor activity | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0060964 | involved_in regulation of miRNA-mediated gene silencing | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0080135 | involved_in regulation of cellular response to stress | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0097009 | involved_in energy homeostasis | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:0098794 | located_in postsynapse | 1 | IEA | Homo_sapiens(9606) | Component |
| GO:1900087 | involved_in positive regulation of G1/S transition of mitotic cell cycle | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:1901215 | involved_in negative regulation of neuron death | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:1905537 | involved_in positive regulation of eukaryotic translation initiation factor 4F complex assembly | 1 | IMP | Homo_sapiens(9606) | Process |
| GO:1905606 | involved_in regulation of presynapse assembly | 1 | IGI | Homo_sapiens(9606) | Process |
| GO:1905612 | involved_in positive regulation of mRNA cap binding | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:1905696 | involved_in regulation of polysome binding | 1 | IMP | Homo_sapiens(9606) | Process |