Welcome to
RBP World!
Functions and Diseases
| RBP Type | Non-canonical_RBPs |
| Diseases | Cancer |
| Drug | N.A. |
| Main interacting RNAs | N.A. |
| Moonlighting functions | N.A. |
| Localizations | Stress granucleP-body |
| BulkPerturb-seq | N.A. |
Description
| Ensembl ID | ENSG00000163220 | Gene ID | 6280 | Accession | 10499 |
| Symbol | S100A9 | Alias | MIF;NIF;P14;CAGB;CFAG;CGLB;L1AG;LIAG;MRP14;60B8AG;MAC387;S100-A9 | Full Name | S100 calcium binding protein A9 |
| Status | Confidence | Length | 3170 bases | Strand | Plus strand |
| Position | 1 : 153357854 - 153361023 | RNA binding domain | N.A. | ||
| Summary | The protein encoded by this gene is a member of the S100 family of proteins containing 2 EF-hand calcium-binding motifs. S100 proteins are localized in the cytoplasm and/or nucleus of a wide range of cells, and involved in the regulation of a number of cellular processes such as cell cycle progression and differentiation. S100 genes include at least 13 members which are located as a cluster on chromosome 1q21. This protein may function in the inhibition of casein kinase and altered expression of this protein is associated with the disease cystic fibrosis. This antimicrobial protein exhibits antifungal and antibacterial activity. [provided by RefSeq, Nov 2014] | ||||
RNA binding domains (RBDs)
| Protein | Domain | Pfam ID | E-value | Domain number |
|---|
RNA binding proteomes (RBPomes)
| Pubmed ID | Full Name | Cell | Author | Time | Doi |
|---|
Literatures on RNA binding capacity
| Pubmed ID | Title | Author | Time | Journal |
|---|---|---|---|---|
| 33335790 | Long Non-Coding RNA Hotairm1 Promotes S100A9 Support of MDSC Expansion during Sepsis. | Tuqa Alkhateeb | 2020-01-01 | Journal of clinical & cellular immunology |
Transcripts
| Name | Transcript ID | bp | Protein | Translation ID |
|---|---|---|---|---|
| S100A9-201 | ENST00000368738 | 573 | 114aa | ENSP00000357727 |
Phenotypes
| ensgID | Trait | pValue | Pubmed ID |
|---|---|---|---|
| 51131 | Child Development Disorders, Pervasive | 4.3329000E-005 | - |
GWAS
| ensgID | SNP | Chromosome | Position | Trait | PubmedID | Or or BEAT | EFO ID |
|---|
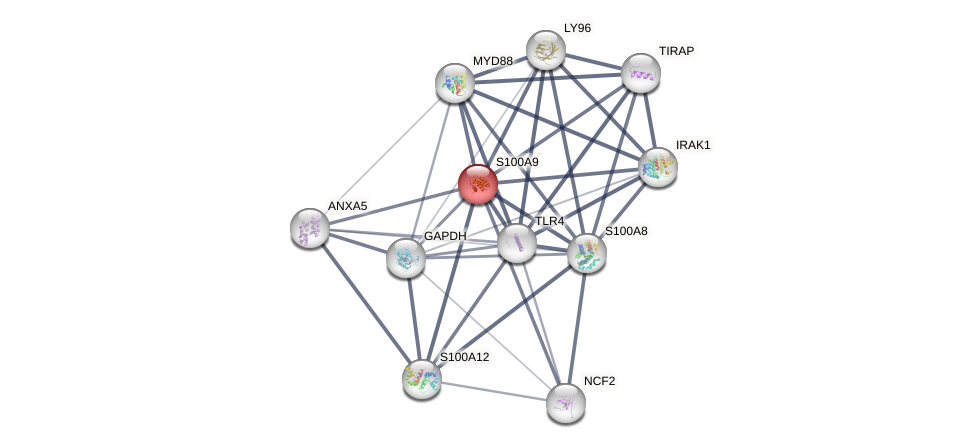
Protein-Protein Interaction (PPI)
Paralogs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| ENSG00000163220 | homo_sapiens | ENSP00000507299 | 54.0816 | 33.6735 | homo_sapiens | ENSP00000357727 | 46.4912 | 28.9474 |
| ENSG00000163220 | homo_sapiens | ENSP00000357702 | 45.5446 | 27.7228 | homo_sapiens | ENSP00000357727 | 40.3509 | 24.5614 |
| ENSG00000163220 | homo_sapiens | ENSP00000369547 | 60.7595 | 31.6456 | homo_sapiens | ENSP00000357727 | 42.1053 | 21.9298 |
| ENSG00000163220 | homo_sapiens | ENSP00000357722 | 63.4409 | 27.957 | homo_sapiens | ENSP00000357727 | 51.7544 | 22.807 |
| ENSG00000163220 | homo_sapiens | ENSP00000341442 | 29.2517 | 17.0068 | homo_sapiens | ENSP00000357727 | 37.7193 | 21.9298 |
| ENSG00000163220 | homo_sapiens | ENSP00000357695 | 51.4563 | 23.301 | homo_sapiens | ENSP00000357727 | 46.4912 | 21.0526 |
| ENSG00000163220 | homo_sapiens | ENSP00000291700 | 61.9565 | 38.0435 | homo_sapiens | ENSP00000357727 | 50 | 30.7018 |
| ENSG00000163220 | homo_sapiens | ENSP00000340463 | 43.2692 | 25.9615 | homo_sapiens | ENSP00000357727 | 39.4737 | 23.6842 |
| ENSG00000163220 | homo_sapiens | ENSP00000357726 | 70.6522 | 46.7391 | homo_sapiens | ENSP00000357727 | 57.0175 | 37.7193 |
| ENSG00000163220 | homo_sapiens | ENSP00000320430 | 59.596 | 33.3333 | homo_sapiens | ENSP00000357727 | 51.7544 | 28.9474 |
| ENSG00000163220 | homo_sapiens | ENSP00000357697 | 58.1633 | 30.6122 | homo_sapiens | ENSP00000357727 | 50 | 26.3158 |
| ENSG00000163220 | homo_sapiens | ENSP00000357801 | 54.6392 | 29.8969 | homo_sapiens | ENSP00000357727 | 46.4912 | 25.4386 |
| ENSG00000163220 | homo_sapiens | ENSP00000357705 | 56.4356 | 27.7228 | homo_sapiens | ENSP00000357727 | 50 | 24.5614 |
| ENSG00000163220 | homo_sapiens | ENSP00000357712 | 50.495 | 27.7228 | homo_sapiens | ENSP00000357727 | 44.7368 | 24.5614 |
| ENSG00000163220 | homo_sapiens | ENSP00000357718 | 50.495 | 26.7327 | homo_sapiens | ENSP00000357727 | 44.7368 | 23.6842 |
| ENSG00000163220 | homo_sapiens | ENSP00000271835 | 11.9192 | 6.66667 | homo_sapiens | ENSP00000357727 | 51.7544 | 28.9474 |
| ENSG00000163220 | homo_sapiens | ENSP00000357706 | 61.9565 | 32.6087 | homo_sapiens | ENSP00000357727 | 50 | 26.3158 |
| ENSG00000163220 | homo_sapiens | ENSP00000292169 | 59.5745 | 35.1064 | homo_sapiens | ENSP00000357727 | 49.1228 | 28.9474 |
| ENSG00000163220 | homo_sapiens | ENSP00000296370 | 60 | 36.8421 | homo_sapiens | ENSP00000357727 | 50 | 30.7018 |
| ENSG00000163220 | homo_sapiens | ENSP00000271638 | 53.3333 | 33.3333 | homo_sapiens | ENSP00000357727 | 49.1228 | 30.7018 |
| ENSG00000163220 | homo_sapiens | ENSP00000357708 | 61.1111 | 31.1111 | homo_sapiens | ENSP00000357727 | 48.2456 | 24.5614 |
Orthologs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 96.4912 | 95.614 | nomascus_leucogenys | ENSNLEP00000012643 | 98.2143 | 97.3214 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 100 | 99.1228 | pan_paniscus | ENSPPAP00000012904 | 100 | 99.1228 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 98.2456 | 97.3684 | pongo_abelii | ENSPPYP00000000953 | 98.2456 | 97.3684 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 100 | 99.1228 | pan_troglodytes | ENSPTRP00000002268 | 100 | 99.1228 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 100 | 100 | gorilla_gorilla | ENSGGOP00000000844 | 100 | 100 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 69.2982 | 51.7544 | ornithorhynchus_anatinus | ENSOANP00000033943 | 64.2276 | 47.9675 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 59.6491 | dasypus_novemcinctus | ENSDNOP00000012977 | 68.75 | 53.125 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.0702 | 60.5263 | dasypus_novemcinctus | ENSDNOP00000031977 | 68.4615 | 53.0769 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 61.4035 | erinaceus_europaeus | ENSEEUP00000004568 | 82.1429 | 62.5 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 42.9825 | 33.3333 | echinops_telfairi | ENSETEP00000007865 | 79.0323 | 61.2903 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 63.1579 | callithrix_jacchus | ENSCJAP00000093953 | 89.1304 | 78.2609 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 88.5965 | cercocebus_atys | ENSCATP00000037753 | 93.8596 | 88.5965 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 88.5965 | macaca_fascicularis | ENSMFAP00000027452 | 93.8596 | 88.5965 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 88.5965 | macaca_mulatta | ENSMMUP00000009023 | 93.8596 | 88.5965 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 88.5965 | macaca_nemestrina | ENSMNEP00000020177 | 93.8596 | 88.5965 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 88.5965 | papio_anubis | ENSPANP00000049485 | 93.8596 | 88.5965 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 88.5965 | mandrillus_leucophaeus | ENSMLEP00000004853 | 93.8596 | 88.5965 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 75.4386 | 58.7719 | tursiops_truncatus | ENSTTRP00000000015 | 76.7857 | 59.8214 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.9474 | 66.6667 | vulpes_vulpes | ENSVVUP00000039621 | 48.913 | 41.3043 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 57.8947 | mus_spretus | MGP_SPRETEiJ_P0061252 | 72.5664 | 58.4071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 82.4561 | 70.1754 | equus_asinus | ENSEASP00005001236 | 61.4379 | 52.2876 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 67.5439 | equus_asinus | ENSEASP00005037914 | 62.1622 | 52.027 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 67.5439 | equus_asinus | ENSEASP00005040644 | 62.1622 | 52.027 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 62.2807 | capra_hircus | ENSCHIP00000016670 | 58.2781 | 47.0199 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.9474 | 66.6667 | loxodonta_africana | ENSLAFP00000024279 | 79.646 | 67.2566 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 61.4035 | panthera_leo | ENSPLOP00000014467 | 64.4444 | 51.8519 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 64.9123 | ailuropoda_melanoleuca | ENSAMEP00000012878 | 78.5714 | 66.0714 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 63.1579 | 51.7544 | ailuropoda_melanoleuca | ENSAMEP00000034168 | 66.6667 | 54.6296 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.0702 | 69.2982 | bos_indicus_hybrid | ENSBIXP00005028500 | 57.0513 | 50.641 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 66.6667 | oryctolagus_cuniculus | ENSOCUP00000007456 | 69.697 | 57.5758 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 79.8246 | 66.6667 | equus_caballus | ENSECAP00000073848 | 51.7045 | 43.1818 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 82.4561 | 70.1754 | equus_caballus | ENSECAP00000077855 | 36.8627 | 31.3725 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 66.6667 | equus_caballus | ENSECAP00000029371 | 56.7901 | 46.9136 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 64.0351 | canis_lupus_familiaris | ENSCAFP00845003183 | 47.8261 | 39.6739 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 61.4035 | panthera_pardus | ENSPPRP00000010241 | 64.4444 | 51.8519 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 83.3333 | 68.4211 | marmota_marmota_marmota | ENSMMMP00000020557 | 83.3333 | 68.4211 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 59.6491 | balaenoptera_musculus | ENSBMSP00010026180 | 59.4203 | 49.2754 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.0526 | 63.1579 | cavia_porcellus | ENSCPOP00000001521 | 67.5 | 60 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 75.4386 | 61.4035 | felis_catus | ENSFCAP00000002948 | 63.7037 | 51.8519 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 58.7719 | 50.8772 | ursus_americanus | ENSUAMP00000011673 | 77.0115 | 66.6667 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 64.0351 | ursus_americanus | ENSUAMP00000003947 | 67.6923 | 56.1538 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.0702 | 68.4211 | bison_bison_bison | ENSBBBP00000020436 | 73.5537 | 64.4628 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 58.7719 | monodon_monoceros | ENSMMNP00015002580 | 67.9688 | 52.3438 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 79.8246 | 64.9123 | sus_scrofa | ENSSSCP00000029038 | 40.991 | 33.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 82.4561 | 71.9298 | microcebus_murinus | ENSMICP00000025254 | 61.039 | 53.2468 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 59.6491 | octodon_degus | ENSODEP00000013144 | 72.5664 | 60.177 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 64.0351 | jaculus_jaculus | ENSJJAP00000001804 | 84.4037 | 66.9725 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 60.5263 | ovis_aries_rambouillet | ENSOARP00020000837 | 59.0604 | 46.3087 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 64.0351 | ursus_maritimus | ENSUMAP00000028538 | 67.6923 | 56.1538 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 81.5789 | 65.7895 | ochotona_princeps | ENSOPRP00000009233 | 79.4872 | 64.1026 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 58.7719 | delphinapterus_leucas | ENSDLEP00000013643 | 55.7692 | 42.9487 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 74.5614 | 57.8947 | physeter_catodon | ENSPCTP00005023199 | 59.8592 | 46.4789 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.0526 | 57.8947 | mus_caroli | MGP_CAROLIEiJ_P0058896 | 71.6814 | 58.4071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 57.8947 | mus_musculus | ENSMUSP00000112843 | 72.5664 | 58.4071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 58.7719 | 42.1053 | mus_musculus | ENSMUSP00000137232 | 57.265 | 41.0256 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.0526 | 53.5088 | dipodomys_ordii | ENSDORP00000003205 | 72.3214 | 54.4643 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 74.5614 | 61.4035 | mustela_putorius_furo | ENSMPUP00000008321 | 44.9735 | 37.037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 83.3333 | 72.807 | aotus_nancymaae | ENSANAP00000000002 | 72.5191 | 63.3588 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 81.5789 | 69.2982 | saimiri_boliviensis_boliviensis | ENSSBOP00000030195 | 72.093 | 61.2403 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 93.8596 | 86.8421 | chlorocebus_sabaeus | ENSCSAP00000016185 | 93.8596 | 86.8421 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 75.4386 | 59.6491 | tupaia_belangeri | ENSTBEP00000010754 | 74.1379 | 58.6207 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 61.4035 | mesocricetus_auratus | ENSMAUP00000004544 | 77.8761 | 61.9469 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 58.7719 | phocoena_sinus | ENSPSNP00000001582 | 79.8165 | 61.4679 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.0702 | 62.2807 | camelus_dromedarius | ENSCDRP00005023762 | 59.7315 | 47.651 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 64.0351 | carlito_syrichta | ENSTSYP00000030358 | 69.1729 | 54.8872 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 62.2807 | panthera_tigris_altaica | ENSPTIP00000026473 | 64.4444 | 52.5926 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 61.4035 | rattus_norvegicus | ENSRNOP00000015351 | 61.6541 | 52.6316 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 79.8246 | 71.0526 | prolemur_simus | ENSPSMP00000029668 | 77.7778 | 69.2308 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 73.6842 | 55.2632 | microtus_ochrogaster | ENSMOCP00000010160 | 73.6842 | 55.2632 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 62.2807 | 38.5965 | microtus_ochrogaster | ENSMOCP00000001691 | 68.2692 | 42.3077 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 64.9123 | 45.614 | peromyscus_maniculatus_bairdii | ENSPEMP00000030403 | 52.4823 | 36.8794 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 72.807 | 59.6491 | peromyscus_maniculatus_bairdii | ENSPEMP00000019308 | 73.4513 | 60.177 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.0702 | 68.4211 | bos_mutus | ENSBMUP00000029040 | 46.8421 | 41.0526 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 73.6842 | 57.8947 | mus_spicilegus | ENSMSIP00000001581 | 74.3363 | 58.4071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 58.7719 | 40.3509 | cricetulus_griseus_chok1gshd | ENSCGRP00001008535 | 54.0323 | 37.0968 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 72.807 | 57.8947 | cricetulus_griseus_chok1gshd | ENSCGRP00001010282 | 73.4513 | 58.4071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 75.4386 | 61.4035 | pteropus_vampyrus | ENSPVAP00000013142 | 72.2689 | 58.8235 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 74.5614 | 61.4035 | myotis_lucifugus | ENSMLUP00000020322 | 56.2914 | 46.3576 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 73.6842 | 59.6491 | rhinolophus_ferrumequinum | ENSRFEP00010021696 | 57.931 | 46.8966 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 62.2807 | neovison_vison | ENSNVIP00000018949 | 66.9231 | 54.6154 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 68.4211 | bos_taurus | ENSBTAP00000067294 | 56.4103 | 50 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 76.3158 | 61.4035 | heterocephalus_glaber_female | ENSHGLP00000038946 | 42.6471 | 34.3137 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 94.7368 | 86.8421 | rhinopithecus_bieti | ENSRBIP00000008437 | 94.7368 | 86.8421 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 64.0351 | canis_lupus_dingo | ENSCAFP00020012872 | 47.8261 | 39.6739 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.9474 | 61.4035 | sciurus_vulgaris | ENSSVLP00005012438 | 80.3571 | 62.5 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 79.8246 | 68.4211 | procavia_capensis | ENSPCAP00000007718 | 79.1304 | 67.8261 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 65.7895 | nannospalax_galili | ENSNGAP00000020406 | 78.5714 | 66.9643 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 77.193 | 67.5439 | bos_grunniens | ENSBGRP00000007988 | 80.7339 | 70.6422 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 69.2982 | 62.2807 | chinchilla_lanigera | ENSCLAP00000004722 | 71.1712 | 63.964 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.9474 | 66.6667 | cervus_hanglu_yarkandensis | ENSCHYP00000027926 | 61.6438 | 52.0548 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 81.5789 | 66.6667 | catagonus_wagneri | ENSCWAP00000003611 | 65.035 | 53.1469 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 81.5789 | 63.1579 | ictidomys_tridecemlineatus | ENSSTOP00000016084 | 80.8696 | 62.6087 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 84.2105 | 71.9298 | otolemur_garnettii | ENSOGAP00000021064 | 83.4783 | 71.3043 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 85.0877 | 73.6842 | cebus_imitator | ENSCCAP00000021817 | 88.1818 | 76.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 81.5789 | 65.7895 | urocitellus_parryii | ENSUPAP00010019916 | 81.5789 | 65.7895 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 78.0702 | 65.7895 | moschus_moschiferus | ENSMMSP00000001552 | 62.6761 | 52.8169 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 94.7368 | 86.8421 | rhinopithecus_roxellana | ENSRROP00000007870 | 94.7368 | 86.8421 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 71.9298 | 57.8947 | mus_pahari | MGP_PahariEiJ_P0069848 | 72.5664 | 58.4071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 80.7018 | 70.1754 | propithecus_coquereli | ENSPCOP00000015193 | 59.7403 | 51.9481 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 30.7018 | lepisosteus_oculatus | ENSLOCP00000008622 | 55.3398 | 33.9806 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 21.0526 | lepisosteus_oculatus | ENSLOCP00000008674 | 50.9804 | 23.5294 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 29.8246 | clupea_harengus | ENSCHAP00000031007 | 58.1633 | 34.6939 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 31.5789 | clupea_harengus | ENSCHAP00000039744 | 56.5657 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 46.4912 | 26.3158 | clupea_harengus | ENSCHAP00000037822 | 55.2083 | 31.25 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | danio_rerio | ENSDARP00000115567 | 50 | 30.3279 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 29.8246 | danio_rerio | ENSDARP00000054468 | 50.9804 | 33.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 29.8246 | danio_rerio | ENSDARP00000072444 | 54.955 | 30.6306 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 28.9474 | carassius_auratus | ENSCARP00000037133 | 53.3333 | 31.4286 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 28.9474 | carassius_auratus | ENSCARP00000046261 | 53.3333 | 31.4286 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 31.5789 | carassius_auratus | ENSCARP00000025961 | 51.4286 | 34.2857 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 28.9474 | carassius_auratus | ENSCARP00000047178 | 53.4653 | 32.6733 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 29.8246 | carassius_auratus | ENSCARP00000038152 | 52.9412 | 33.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 24.5614 | carassius_auratus | ENSCARP00000025198 | 57.732 | 28.866 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.0702 | carassius_auratus | ENSCARP00000033672 | 39.1026 | 20.5128 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | astyanax_mexicanus | ENSAMXP00000020513 | 48.4127 | 29.3651 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 31.5789 | astyanax_mexicanus | ENSAMXP00000046346 | 60.396 | 35.6436 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 27.193 | astyanax_mexicanus | ENSAMXP00000052348 | 37.1069 | 19.4969 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 32.4561 | ictalurus_punctatus | ENSIPUP00000010089 | 42.7586 | 25.5172 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 30.7018 | ictalurus_punctatus | ENSIPUP00000011251 | 59.4059 | 34.6535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | ictalurus_punctatus | ENSIPUP00000005660 | 55.0459 | 30.2752 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 30.7018 | esox_lucius | ENSELUP00000049881 | 50.4065 | 28.4553 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 31.5789 | esox_lucius | ENSELUP00000068084 | 59.596 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 28.0702 | esox_lucius | ENSELUP00000072348 | 53.271 | 29.9065 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 29.8246 | electrophorus_electricus | ENSEEEP00000012970 | 54.2857 | 32.381 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 31.5789 | electrophorus_electricus | ENSEEEP00000013321 | 60.396 | 35.6436 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 24.5614 | electrophorus_electricus | ENSEEEP00000023160 | 57.2917 | 29.1667 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 31.5789 | oncorhynchus_kisutch | ENSOKIP00005064078 | 52.9412 | 30.2521 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 32.4561 | oncorhynchus_kisutch | ENSOKIP00005049402 | 51.2195 | 30.0813 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 32.4561 | oncorhynchus_kisutch | ENSOKIP00005098713 | 26.5823 | 15.6118 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 30.7018 | oncorhynchus_kisutch | ENSOKIP00005063008 | 59.596 | 35.3535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | oncorhynchus_kisutch | ENSOKIP00005000280 | 60.396 | 36.6337 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 31.5789 | oncorhynchus_kisutch | ENSOKIP00005049075 | 40.5405 | 24.3243 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | oncorhynchus_kisutch | ENSOKIP00005100281 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | oncorhynchus_kisutch | ENSOKIP00005029712 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 26.3158 | oncorhynchus_kisutch | ENSOKIP00005062955 | 35.1515 | 18.1818 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 32.4561 | oncorhynchus_mykiss | ENSOMYP00000011858 | 50 | 29.3651 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 32.4561 | oncorhynchus_mykiss | ENSOMYP00000109487 | 62.5 | 38.5417 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 33.3333 | oncorhynchus_mykiss | ENSOMYP00000006954 | 33.8983 | 21.4689 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | oncorhynchus_mykiss | ENSOMYP00000139195 | 15.099 | 9.15842 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 30.7018 | oncorhynchus_mykiss | ENSOMYP00000075455 | 59.596 | 35.3535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 32.4561 | oncorhynchus_mykiss | ENSOMYP00000091424 | 44.9275 | 26.8116 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 25.4386 | oncorhynchus_mykiss | ENSOMYP00000112176 | 61.9565 | 31.5217 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | oncorhynchus_mykiss | ENSOMYP00000117475 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | oncorhynchus_mykiss | ENSOMYP00000000231 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 31.5789 | salmo_salar | ENSSSAP00000011588 | 50.8064 | 29.0323 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 32.4561 | salmo_salar | ENSSSAP00000146697 | 51.2195 | 30.0813 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 31.5789 | salmo_salar | ENSSSAP00000169779 | 44.1176 | 26.4706 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | salmo_salar | ENSSSAP00000040712 | 60.396 | 36.6337 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 32.4561 | salmo_salar | ENSSSAP00000172868 | 48.062 | 28.6822 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 31.5789 | salmo_salar | ENSSSAP00000050364 | 41.5493 | 25.3521 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | salmo_salar | ENSSSAP00000076269 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | salmo_salar | ENSSSAP00000028172 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 32.4561 | salmo_trutta | ENSSTUP00000093956 | 51.2195 | 30.0813 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 31.5789 | salmo_trutta | ENSSTUP00000078161 | 50.8064 | 29.0323 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | salmo_trutta | ENSSTUP00000087292 | 61.6162 | 35.3535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 31.5789 | salmo_trutta | ENSSTUP00000094521 | 60.6061 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 31.5789 | salmo_trutta | ENSSTUP00000096680 | 58.4158 | 35.6436 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 30.7018 | salmo_trutta | ENSSTUP00000053418 | 48.7805 | 28.4553 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | salmo_trutta | ENSSTUP00000109619 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 40.3509 | 22.807 | salmo_trutta | ENSSTUP00000109636 | 47.9167 | 27.0833 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 29.8246 | gadus_morhua | ENSGMOP00000011657 | 59.1398 | 36.5591 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 29.8246 | gadus_morhua | ENSGMOP00000039183 | 56.4356 | 33.6634 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.9474 | gadus_morhua | ENSGMOP00000018647 | 33.3333 | 18.0328 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 35.0877 | fundulus_heteroclitus | ENSFHEP00000010342 | 57.1429 | 35.7143 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 31.5789 | fundulus_heteroclitus | ENSFHEP00000028017 | 55 | 36 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | fundulus_heteroclitus | ENSFHEP00000033400 | 56.0748 | 29.9065 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 57.0175 | 35.9649 | poecilia_reticulata | ENSPREP00000029135 | 58.0357 | 36.6071 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | poecilia_reticulata | ENSPREP00000025184 | 56.0748 | 29.9065 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 57.0175 | 36.8421 | xiphophorus_maculatus | ENSXMAP00000014843 | 56.0345 | 36.2069 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 46.4912 | 30.7018 | xiphophorus_maculatus | ENSXMAP00000014781 | 53 | 35 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 27.193 | xiphophorus_maculatus | ENSXMAP00000007243 | 55.3398 | 30.0971 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 34.2105 | oryzias_latipes | ENSORLP00000036516 | 43.4483 | 26.8966 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | oryzias_latipes | ENSORLP00000012201 | 56.4815 | 32.4074 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 34.2105 | cyclopterus_lumpus | ENSCLMP00005035380 | 53.4483 | 33.6207 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | cyclopterus_lumpus | ENSCLMP00005024958 | 56.0748 | 30.8411 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 34.2105 | oreochromis_niloticus | ENSONIP00000058449 | 36.8421 | 22.807 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 46.4912 | 25.4386 | oreochromis_niloticus | ENSONIP00000032624 | 38.4058 | 21.0145 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 27.193 | oreochromis_niloticus | ENSONIP00000007928 | 60.2151 | 33.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 34.2105 | haplochromis_burtoni | ENSHBUP00000018709 | 53.3898 | 33.0508 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 25.4386 | haplochromis_burtoni | ENSHBUP00000018552 | 54.1667 | 30.2083 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | haplochromis_burtoni | ENSHBUP00000023416 | 57.0093 | 32.7103 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 34.2105 | astatotilapia_calliptera | ENSACLP00000015770 | 37.5 | 23.2143 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 25.4386 | astatotilapia_calliptera | ENSACLP00000014601 | 54.7368 | 30.5263 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | astatotilapia_calliptera | ENSACLP00000027616 | 57.0093 | 32.7103 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 33.3333 | sparus_aurata | ENSSAUP00010016074 | 50.8475 | 32.2034 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 32.4561 | sparus_aurata | ENSSAUP00010015528 | 36.1963 | 22.6994 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 35.0877 | sparus_aurata | ENSSAUP00010035228 | 35.503 | 23.6686 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | sparus_aurata | ENSSAUP00010046326 | 25.1046 | 13.8075 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 34.2105 | lates_calcarifer | ENSLCAP00010048401 | 53.7815 | 32.7731 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 33.3333 | lates_calcarifer | ENSLCAP00010048366 | 56 | 38 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | lates_calcarifer | ENSLCAP00010054665 | 42.2535 | 23.2394 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 24.5614 | xenopus_tropicalis | ENSXETP00000018253 | 24.8848 | 12.9032 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 32.4561 | neolamprologus_brichardi | ENSNBRP00000021329 | 56.0748 | 34.5794 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 25.4386 | neolamprologus_brichardi | ENSNBRP00000021809 | 54.1667 | 30.2083 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 31.5789 | neolamprologus_brichardi | ENSNBRP00000006713 | 51.3274 | 31.8584 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 46.4912 | 24.5614 | erpetoichthys_calabaricus | ENSECRP00000003464 | 46.4912 | 24.5614 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 57.0175 | 35.0877 | kryptolebias_marmoratus | ENSKMAP00000015076 | 54.6218 | 33.6134 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 29.8246 | kryptolebias_marmoratus | ENSKMAP00000015302 | 55 | 34 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 29.8246 | kryptolebias_marmoratus | ENSKMAP00000003514 | 55.5556 | 31.4815 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 33.3333 | oryzias_javanicus | ENSOJAP00000011971 | 55.3571 | 33.9286 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 29.8246 | oryzias_javanicus | ENSOJAP00000011559 | 57.2917 | 35.4167 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | oryzias_javanicus | ENSOJAP00000035273 | 57.0093 | 32.7103 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 34.2105 | amphilophus_citrinellus | ENSACIP00000005032 | 54.386 | 34.2105 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 44.7368 | 26.3158 | amphilophus_citrinellus | ENSACIP00000002311 | 53.125 | 31.25 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 29.8246 | amphilophus_citrinellus | ENSACIP00000013575 | 56.7308 | 32.6923 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 34.2105 | dicentrarchus_labrax | ENSDLAP00005031210 | 55.3571 | 34.8214 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 29.8246 | dicentrarchus_labrax | ENSDLAP00005079063 | 56.701 | 35.0515 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | dicentrarchus_labrax | ENSDLAP00005068331 | 42.8571 | 23.5714 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | amphiprion_percula | ENSAPEP00000011539 | 56.4815 | 34.2593 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 31.5789 | amphiprion_percula | ENSAPEP00000005448 | 56.1224 | 36.7347 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 32.4561 | amphiprion_percula | ENSAPEP00000011858 | 58 | 37 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 28.9474 | amphiprion_percula | ENSAPEP00000004850 | 54.6296 | 30.5556 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 34.2105 | oryzias_sinensis | ENSOSIP00000003741 | 56.25 | 34.8214 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 28.9474 | oryzias_sinensis | ENSOSIP00000003160 | 31.4286 | 18.8571 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 30.7018 | oryzias_sinensis | ENSOSIP00000024811 | 43.662 | 24.6479 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 33.3333 | seriola_dumerili | ENSSDUP00000024787 | 61.2245 | 38.7755 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 29.8246 | seriola_dumerili | ENSSDUP00000008052 | 58.1633 | 34.6939 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 33.3333 | seriola_dumerili | ENSSDUP00000025050 | 57 | 38 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | seriola_dumerili | ENSSDUP00000007378 | 55.5556 | 30.5556 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 29.8246 | cottoperca_gobio | ENSCGOP00000030780 | 58.3333 | 35.4167 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 31.5789 | cottoperca_gobio | ENSCGOP00000030678 | 55.5556 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | cottoperca_gobio | ENSCGOP00000017925 | 42.8571 | 23.5714 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 32.4561 | gasterosteus_aculeatus | ENSGACP00000005933 | 56.8627 | 36.2745 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 30.7018 | gasterosteus_aculeatus | ENSGACP00000004450 | 56.8627 | 34.3137 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 29.8246 | gasterosteus_aculeatus | ENSGACP00000015839 | 52.9412 | 33.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | gasterosteus_aculeatus | ENSGACP00000016004 | 55.2381 | 29.5238 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 32.4561 | cynoglossus_semilaevis | ENSCSEP00000011456 | 54.8673 | 32.7434 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 29.8246 | cynoglossus_semilaevis | ENSCSEP00000011680 | 48.6957 | 29.5652 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 27.193 | cynoglossus_semilaevis | ENSCSEP00000003340 | 37.931 | 21.3793 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 26.3158 | cynoglossus_semilaevis | ENSCSEP00000003158 | 56.1224 | 30.6122 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 35.9649 | poecilia_formosa | ENSPFOP00000009736 | 57.6577 | 36.9369 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 31.5789 | poecilia_formosa | ENSPFOP00000009357 | 47.7876 | 31.8584 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | poecilia_formosa | ENSPFOP00000015246 | 56.0748 | 29.9065 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 32.4561 | myripristis_murdjan | ENSMMDP00005042081 | 50.8475 | 31.3559 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 29.8246 | myripristis_murdjan | ENSMMDP00005032634 | 57 | 34 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 28.9474 | myripristis_murdjan | ENSMMDP00005041605 | 49.0909 | 30 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 25.4386 | myripristis_murdjan | ENSMMDP00005031606 | 59.1398 | 31.1828 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 34.2105 | sander_lucioperca | ENSSLUP00000028673 | 53.5088 | 34.2105 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 33.3333 | sander_lucioperca | ENSSLUP00000028578 | 52.2523 | 34.2342 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 29.8246 | sander_lucioperca | ENSSLUP00000026687 | 54.5455 | 34.3434 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 25.4386 | sander_lucioperca | ENSSLUP00000025912 | 59.1398 | 31.1828 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | amphiprion_ocellaris | ENSAOCP00000005684 | 56.4815 | 34.2593 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 32.4561 | amphiprion_ocellaris | ENSAOCP00000007600 | 57.1429 | 37.7551 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 32.4561 | amphiprion_ocellaris | ENSAOCP00000027574 | 58 | 37 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 28.9474 | amphiprion_ocellaris | ENSAOCP00000021093 | 54.6296 | 30.5556 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 57.0175 | 31.5789 | hucho_hucho | ENSHHUP00000041660 | 29.2793 | 16.2162 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 31.5789 | hucho_hucho | ENSHHUP00000059948 | 34.8066 | 19.8895 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 31.5789 | hucho_hucho | ENSHHUP00000041880 | 60.6061 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 30.7018 | hucho_hucho | ENSHHUP00000088409 | 43.0556 | 24.3056 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 33.3333 | hucho_hucho | ENSHHUP00000018498 | 33.1579 | 20 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 31.5789 | hucho_hucho | ENSHHUP00000021509 | 59.4059 | 35.6436 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | hucho_hucho | ENSHHUP00000023969 | 60.396 | 34.6535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 27.193 | hucho_hucho | ENSHHUP00000084732 | 24.7934 | 12.8099 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | hucho_hucho | ENSHHUP00000019484 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 30.7018 | betta_splendens | ENSBSLP00000032820 | 58.5859 | 35.3535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 25.4386 | betta_splendens | ENSBSLP00000034713 | 59.1398 | 31.1828 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 32.4561 | pygocentrus_nattereri | ENSPNAP00000002994 | 47.619 | 29.3651 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | pygocentrus_nattereri | ENSPNAP00000011775 | 60.396 | 36.6337 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | pygocentrus_nattereri | ENSPNAP00000026045 | 53.0973 | 28.3186 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 34.2105 | anabas_testudineus | ENSATEP00000013665 | 54.2373 | 33.0508 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 31.5789 | anabas_testudineus | ENSATEP00000013445 | 57 | 36 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | anabas_testudineus | ENSATEP00000060137 | 55.5556 | 30.5556 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 35.0877 | seriola_lalandi_dorsalis | ENSSLDP00000018686 | 52.8926 | 33.0578 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | seriola_lalandi_dorsalis | ENSSLDP00000018707 | 55.5556 | 30.5556 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 54.386 | 34.2105 | labrus_bergylta | ENSLBEP00000035211 | 51.2397 | 32.2314 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 32.4561 | labrus_bergylta | ENSLBEP00000035522 | 57 | 37 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 29.8246 | labrus_bergylta | ENSLBEP00000006895 | 36.1446 | 20.4819 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 30.7018 | pundamilia_nyererei | ENSPNYP00000014053 | 58 | 35 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 25.4386 | pundamilia_nyererei | ENSPNYP00000014338 | 54.1667 | 30.2083 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | pundamilia_nyererei | ENSPNYP00000007026 | 57.0093 | 32.7103 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 35.0877 | mastacembelus_armatus | ENSMAMP00000024826 | 55.6522 | 34.7826 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 27.193 | mastacembelus_armatus | ENSMAMP00000021254 | 37.8378 | 20.9459 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 31.5789 | mastacembelus_armatus | ENSMAMP00000000666 | 53.7037 | 33.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 25.4386 | mastacembelus_armatus | ENSMAMP00000020608 | 59.7826 | 31.5217 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 24.5614 | stegastes_partitus | ENSSPAP00000003398 | 59.1398 | 30.1075 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 35.0877 | stegastes_partitus | ENSSPAP00000029358 | 55.1724 | 34.4828 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 43.8596 | 21.9298 | stegastes_partitus | ENSSPAP00000018365 | 58.1395 | 29.0698 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 33.3333 | stegastes_partitus | ENSSPAP00000003389 | 57 | 38 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 30.7018 | stegastes_partitus | ENSSPAP00000029612 | 55.102 | 35.7143 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 28.9474 | stegastes_partitus | ENSSPAP00000024038 | 56.3107 | 32.0388 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 31.5789 | oncorhynchus_tshawytscha | ENSOTSP00005005100 | 51.2195 | 29.2683 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 57.0175 | 32.4561 | oncorhynchus_tshawytscha | ENSOTSP00005020281 | 32.1782 | 18.3168 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 30.7018 | oncorhynchus_tshawytscha | ENSOTSP00005101225 | 59.596 | 35.3535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 31.5789 | oncorhynchus_tshawytscha | ENSOTSP00005020328 | 51.2195 | 29.2683 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 25.4386 | oncorhynchus_tshawytscha | ENSOTSP00005101221 | 43.1818 | 21.9697 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 27.193 | oncorhynchus_tshawytscha | ENSOTSP00005084674 | 53.7037 | 28.7037 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 34.2105 | acanthochromis_polyacanthus | ENSAPOP00000027122 | 37.6471 | 22.9412 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 31.5789 | acanthochromis_polyacanthus | ENSAPOP00000003981 | 56 | 36 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 32.4561 | acanthochromis_polyacanthus | ENSAPOP00000006958 | 61.2245 | 37.7551 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 25.4386 | acanthochromis_polyacanthus | ENSAPOP00000031991 | 55.6701 | 29.8969 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 30.7018 | nothobranchius_furzeri | ENSNFUP00015023317 | 58 | 35 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 31.5789 | nothobranchius_furzeri | ENSNFUP00015043467 | 35.2201 | 22.6415 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.9474 | nothobranchius_furzeri | ENSNFUP00015025441 | 56.0748 | 30.8411 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 56.1404 | 35.0877 | cyprinodon_variegatus | ENSCVAP00000014796 | 54.7009 | 34.188 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 30.7018 | cyprinodon_variegatus | ENSCVAP00000029662 | 55.102 | 35.7143 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | cyprinodon_variegatus | ENSCVAP00000032878 | 36.1446 | 19.2771 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 24.5614 | denticeps_clupeoides | ENSDCDP00000039852 | 28.5714 | 15.3846 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 30.7018 | denticeps_clupeoides | ENSDCDP00000017084 | 56.1224 | 35.7143 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 24.5614 | denticeps_clupeoides | ENSDCDP00000059689 | 60.6383 | 29.7872 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 35.0877 | larimichthys_crocea | ENSLCRP00005053706 | 54.3103 | 34.4828 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 34.2105 | larimichthys_crocea | ENSLCRP00005053520 | 58.5859 | 39.3939 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | larimichthys_crocea | ENSLCRP00005024889 | 55.5556 | 29.6296 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 34.2105 | takifugu_rubripes | ENSTRUP00000079616 | 56.4815 | 36.1111 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 33.3333 | takifugu_rubripes | ENSTRUP00000073658 | 42.5532 | 26.9504 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 31.5789 | takifugu_rubripes | ENSTRUP00000028045 | 55.5556 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 31.5789 | takifugu_rubripes | ENSTRUP00000070345 | 54.5455 | 36.3636 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.9474 | takifugu_rubripes | ENSTRUP00000084550 | 27.8539 | 15.0685 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 34.2105 | hippocampus_comes | ENSHCOP00000001611 | 54.955 | 35.1351 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50.8772 | 30.7018 | hippocampus_comes | ENSHCOP00000001660 | 58 | 35 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 28.0702 | hippocampus_comes | ENSHCOP00000006402 | 54.6296 | 29.6296 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | cyprinus_carpio_carpio | ENSCCRP00000155002 | 48.8 | 29.6 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 32.4561 | cyprinus_carpio_carpio | ENSCCRP00000109902 | 45.8647 | 27.8195 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 30.7018 | cyprinus_carpio_carpio | ENSCCRP00000127229 | 54.4554 | 34.6535 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 28.9474 | cyprinus_carpio_carpio | ENSCCRP00000142142 | 53.4653 | 32.6733 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 46.4912 | 28.9474 | cyprinus_carpio_carpio | ENSCCRP00000131167 | 56.383 | 35.1064 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.0702 | cyprinus_carpio_carpio | ENSCCRP00000110516 | 43.8849 | 23.0216 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.0702 | cyprinus_carpio_carpio | ENSCCRP00000137347 | 43.5714 | 22.8571 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 34.2105 | poecilia_latipinna | ENSPLAP00000012287 | 55.4545 | 35.4545 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 31.5789 | poecilia_latipinna | ENSPLAP00000029132 | 55.6701 | 37.1134 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 28.0702 | poecilia_latipinna | ENSPLAP00000023674 | 56.0748 | 29.9065 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 33.3333 | sinocyclocheilus_grahami | ENSSGRP00000011995 | 48 | 30.4 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 32.4561 | sinocyclocheilus_grahami | ENSSGRP00000111570 | 51.2821 | 31.6239 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 28.9474 | sinocyclocheilus_grahami | ENSSGRP00000110879 | 53.4653 | 32.6733 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 30.7018 | sinocyclocheilus_grahami | ENSSGRP00000110877 | 51.8519 | 32.4074 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.0702 | sinocyclocheilus_grahami | ENSSGRP00000078644 | 53.9823 | 28.3186 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 24.5614 | sinocyclocheilus_grahami | ENSSGRP00000014189 | 57.732 | 28.866 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 34.2105 | maylandia_zebra | ENSMZEP00005035549 | 53.3898 | 33.0508 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 25.4386 | maylandia_zebra | ENSMZEP00005035525 | 54.1667 | 30.2083 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 30.7018 | maylandia_zebra | ENSMZEP00005030029 | 57.0093 | 32.7103 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 31.5789 | tetraodon_nigroviridis | ENSTNIP00000020706 | 57.8431 | 35.2941 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 31.5789 | tetraodon_nigroviridis | ENSTNIP00000020701 | 56.4356 | 35.6436 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 45.614 | 22.807 | tetraodon_nigroviridis | ENSTNIP00000001625 | 55.914 | 27.957 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 29.8246 | paramormyrops_kingsleyae | ENSPKIP00000015966 | 57.2816 | 33.0097 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 53.5088 | 28.0702 | paramormyrops_kingsleyae | ENSPKIP00000007725 | 54.955 | 28.8288 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 29.8246 | scleropages_formosus | ENSSFOP00015077877 | 50.8621 | 29.3103 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 50 | 24.5614 | scleropages_formosus | ENSSFOP00015033956 | 55.3398 | 27.1845 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 49.1228 | 29.8246 | scleropages_formosus | ENSSFOP00015015722 | 55.4455 | 33.6634 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 28.9474 | scleropages_formosus | ENSSFOP00015033529 | 35 | 18.3333 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 43.8596 | 23.6842 | leptobrachium_leishanense | ENSLLEP00000030966 | 43.8596 | 23.6842 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 46.4912 | 25.4386 | leptobrachium_leishanense | ENSLLEP00000031053 | 21.2851 | 11.6466 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 55.2632 | 35.0877 | scophthalmus_maximus | ENSSMAP00000062453 | 54.3103 | 34.4828 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 30.7018 | scophthalmus_maximus | ENSSMAP00000068368 | 55 | 35 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 47.3684 | 25.4386 | scophthalmus_maximus | ENSSMAP00000046250 | 58.6957 | 31.5217 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 51.7544 | 31.5789 | oryzias_melastigma | ENSOMEP00000005300 | 57.2816 | 34.9515 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 48.2456 | 29.8246 | oryzias_melastigma | ENSOMEP00000025336 | 57.2917 | 35.4167 |
| ENSG00000163220 | homo_sapiens | ENSP00000357727 | 52.6316 | 30.7018 | oryzias_melastigma | ENSOMEP00000005944 | 55.5556 | 32.4074 |
Gene Ontology
| Go ID | Go term | No. evidence | Entries | Species | Category |
|---|---|---|---|---|---|
| GO:0002523 | involved_in leukocyte migration involved in inflammatory response | 2 | IBA,IDA | Homo_sapiens(9606) | Process |
| GO:0002544 | involved_in chronic inflammatory response | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0005509 | enables calcium ion binding | 2 | IBA,TAS | Homo_sapiens(9606) | Function |
| GO:0005515 | enables protein binding | 1 | IPI | Homo_sapiens(9606) | Function |
| GO:0005576 | located_in extracellular region | 2 | HDA,TAS | Homo_sapiens(9606) | Component |
| GO:0005615 | located_in extracellular space | 2 | HDA,IDA | Homo_sapiens(9606) | Component |
| GO:0005634 | located_in nucleus | 1 | HDA | Homo_sapiens(9606) | Component |
| GO:0005737 | is_active_in cytoplasm | 2 | IBA,IDA | Homo_sapiens(9606) | Component |
| GO:0005829 | located_in cytosol | 1 | TAS | Homo_sapiens(9606) | Component |
| GO:0005856 | located_in cytoskeleton | 1 | TAS | Homo_sapiens(9606) | Component |
| GO:0005886 | located_in plasma membrane | 1 | TAS | Homo_sapiens(9606) | Component |
| GO:0006914 | involved_in autophagy | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:0006915 | involved_in apoptotic process | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0006919 | involved_in activation of cysteine-type endopeptidase activity involved in apoptotic process | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:0006954 | involved_in inflammatory response | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0007267 | involved_in cell-cell signaling | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0008017 | enables microtubule binding | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0008270 | enables zinc ion binding | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0010976 | involved_in positive regulation of neuron projection development | 1 | ISS | Homo_sapiens(9606) | Process |
| GO:0014002 | involved_in astrocyte development | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0016209 | enables antioxidant activity | 1 | IEA | Homo_sapiens(9606) | Function |
| GO:0018119 | involved_in peptidyl-cysteine S-nitrosylation | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0030307 | involved_in positive regulation of cell growth | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0030593 | involved_in neutrophil chemotaxis | 2 | IBA,IDA | Homo_sapiens(9606) | Process |
| GO:0032119 | involved_in sequestering of zinc ion | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0032496 | involved_in response to lipopolysaccharide | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0034774 | located_in secretory granule lumen | 1 | TAS | Homo_sapiens(9606) | Component |
| GO:0035425 | involved_in autocrine signaling | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0035606 | involved_in peptidyl-cysteine S-trans-nitrosylation | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:0035662 | enables Toll-like receptor 4 binding | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0035821 | involved_in modulation of process of another organism | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:0042742 | involved_in defense response to bacterium | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0043542 | involved_in endothelial cell migration | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0045087 | involved_in innate immune response | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:0045113 | involved_in regulation of integrin biosynthetic process | 1 | IBA | Homo_sapiens(9606) | Process |
| GO:0048306 | enables calcium-dependent protein binding | 1 | IBA | Homo_sapiens(9606) | Function |
| GO:0050544 | enables arachidonic acid binding | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0050729 | involved_in positive regulation of inflammatory response | 1 | IDA | Homo_sapiens(9606) | Process |
| GO:0050786 | enables RAGE receptor binding | 1 | TAS | Homo_sapiens(9606) | Function |
| GO:0050832 | involved_in defense response to fungus | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0051092 | involved_in positive regulation of NF-kappaB transcription factor activity | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0051493 | involved_in regulation of cytoskeleton organization | 1 | TAS | Homo_sapiens(9606) | Process |
| GO:0061844 | involved_in antimicrobial humoral immune response mediated by antimicrobial peptide | 2 | IBA,IDA | Homo_sapiens(9606) | Process |
| GO:0062023 | located_in collagen-containing extracellular matrix | 1 | HDA | Homo_sapiens(9606) | Component |
| GO:0070062 | located_in extracellular exosome | 1 | HDA | Homo_sapiens(9606) | Component |
| GO:0070488 | involved_in neutrophil aggregation | 2 | IBA,IDA | Homo_sapiens(9606) | Process |
| GO:0098869 | involved_in cellular oxidant detoxification | 1 | IEA | Homo_sapiens(9606) | Process |
| GO:2001244 | involved_in positive regulation of intrinsic apoptotic signaling pathway | 1 | IDA | Homo_sapiens(9606) | Process |