RBP Type
Non-canonical_RBPs
Diseases
Dengue
Drug
Main interacting RNAs
ncRNA
Moonlighting functions
TF
Localizations
N.A.
BulkPerturb-seq
N.A.
Ensembl ID ENSG00000130816 Gene ID
1786 Accession
2976
Symbol
DNMT1
Alias
AIM;DNMT;MCMT;CXXC9;HSN1E;ADCADN;m.HsaI
Full Name
DNA methyltransferase 1
Status
Confidence
Length
97941 bases
Strand
Minus strand
Position
19 : 10133346 - 10231286
RNA binding domain
N.A.
Summary
This gene encodes an enzyme that transfers methyl groups to cytosine nucleotides of genomic DNA. This protein is the major enzyme responsible for maintaining methylation patterns following DNA replication and shows a preference for hemi-methylated DNA. Methylation of DNA is an important component of mammalian epigenetic gene regulation. Aberrant methylation patterns are found in human tumors and associated with developmental abnormalities. Variation in this gene has been associated with cerebellar ataxia, deafness, and narcolepsy, and neuropathy, hereditary sensory, type IE. Alternative splicing results in multiple transcript variants. [provided by RefSeq, Jan 2016]
RNA binding domains (RBDs)
Protein
Domain
Pfam ID
E-value
Domain number
RNA binding proteomes (RBPomes)
Pubmed ID
Full Name
Cell
Author
Time
Doi
Literatures on RNA binding capacity
Pubmed ID
Title
Author
Time
Journal
36574982 DNMT1 inhibition by pUG-fold quadruplex RNA.
I Linnea Jansson-Fritzberg
2023-03-01
RNA (New York, N.Y.)
19388845 DNA Methyltransferase protein synthesis is reduced in CXXC finger protein 1-deficient embryonic stem cells.
S Jill Butler
2009-05-01
DNA and cell biology
21507353 Dnmt1 structure and function.
M Željko Svedružić
2011-01-01
Progress in molecular biology and translational science
25990724 Small RNA-mediated DNA (cytosine-5) methyltransferase 1 inhibition leads to aberrant DNA methylation.
Guoqiang Zhang
2015-07-13
Nucleic acids research
19048127 A method for purification, identification and validation of DNMT1 mRNA binding proteins.
Alexander Unterberger
2008-01-01
Biological procedures online
31698782 Emerging Roles of Long Non-Coding RNAs as Drivers of Brain Evolution.
Geraldine Zimmer-Bensch
2019-11-06
Cells
34908111 Circular RNAs in stem cells: from basic research to clinical implications.
Hui-Juan Lu
2022-01-28
Bioscience reports
31485661 Circular RNAs: A novel target among non‑coding RNAs with potential roles in malignant tumors (Review).
Weisong Zhao
2019-10-01
Molecular medicine reports
25863539 The long non-coding RNA HNF1A-AS1 regulates proliferation and metastasis in lung adenocarcinoma.
Ying Wu
2015-04-20
Oncotarget
32718982 Functional annotation of human long noncoding RNAs via molecular phenotyping.
A Jordan Ramilowski
2020-07-01
Genome research
29416775 The DNA methyl-transferase protein DNMT1 enhances tumor-promoting properties of breast stromal fibroblasts.
A Layla Al-Kharashi
2018-01-05
Oncotarget
36725842 LncRNA TINCR impairs the efficacy of immunotherapy against breast cancer by recruiting DNMT1 and downregulating MiR-199a-5p via the STAT1-TINCR-USP20-PD-L1 axis.
Qin Wang
2023-02-01
Cell death & disease
30108179 Human proteins that interact with RNA/DNA hybrids.
X Isabel Wang
2018-09-01
Genome research
Name
Transcript ID
bp
Protein
Translation ID
DNMT1-207
ENST00000586667
3259
No protein
-
DNMT1-206
ENST00000586588
3094
No protein
-
DNMT1-227
ENST00000592705
5215
89aa
ENSP00000466657
DNMT1-202
ENST00000359526
5274
1632aa
ENSP00000352516
DNMT1-201
ENST00000340748
5408
1616aa
ENSP00000345739
DNMT1-224
ENST00000591798
521
No protein
-
DNMT1-243
ENST00000678107
2204
No protein
-
DNMT1-219
ENST00000589351
4395
No protein
-
DNMT1-249
ENST00000678957
2873
No protein
-
DNMT1-250
ENST00000679100
3348
No protein
-
DNMT1-246
ENST00000678694
4482
No protein
-
DNMT1-240
ENST00000677783
6469
No protein
-
DNMT1-232
ENST00000676868
5845
No protein
-
DNMT1-242
ENST00000678024
6142
No protein
-
DNMT1-229
ENST00000676604
4821
No protein
-
DNMT1-237
ENST00000677616
5101
1150aa
ENSP00000503055
DNMT1-239
ENST00000677685
4974
184aa
ENSP00000503407
DNMT1-233
ENST00000677013
5175
89aa
ENSP00000503135
DNMT1-236
ENST00000677250
5127
263aa
ENSP00000502894
DNMT1-231
ENST00000676820
6055
No protein
-
DNMT1-238
ENST00000677634
5165
1114aa
ENSP00000504246
DNMT1-251
ENST00000679103
5126
1577aa
ENSP00000503151
DNMT1-234
ENST00000677038
1756
No protein
-
DNMT1-245
ENST00000678647
3302
No protein
-
DNMT1-248
ENST00000678851
3143
No protein
-
DNMT1-230
ENST00000676610
7157
1613aa
ENSP00000504236
DNMT1-214
ENST00000588913
1775
389aa
ENSP00000467125
DNMT1-247
ENST00000678804
5728
1658aa
ENSP00000503853
DNMT1-252
ENST00000679313
5122
1580aa
ENSP00000504512
DNMT1-235
ENST00000677135
653
No protein
-
DNMT1-241
ENST00000677946
5665
1655aa
ENSP00000504202
DNMT1-244
ENST00000678239
2117
No protein
-
DNMT1-217
ENST00000589294
575
No protein
-
DNMT1-211
ENST00000587197
809
No protein
-
DNMT1-220
ENST00000589538
559
No protein
-
DNMT1-228
ENST00000593049
534
No protein
-
DNMT1-212
ENST00000587604
513
No protein
-
DNMT1-216
ENST00000589091
563
No protein
-
DNMT1-222
ENST00000591239
550
No protein
-
DNMT1-208
ENST00000586799
672
55aa
ENSP00000467260
DNMT1-203
ENST00000585843
911
No protein
-
DNMT1-218
ENST00000589349
603
No protein
-
DNMT1-223
ENST00000591764
341
No protein
-
DNMT1-204
ENST00000585920
529
No protein
-
DNMT1-210
ENST00000586988
711
40aa
ENSP00000464958
DNMT1-215
ENST00000588952
841
119aa
ENSP00000467050
DNMT1-213
ENST00000588118
973
255aa
ENSP00000465223
DNMT1-226
ENST00000592342
636
94aa
ENSP00000465993
DNMT1-225
ENST00000592054
591
71aa
ENSP00000468359
DNMT1-221
ENST00000590619
231
77aa
ENSP00000468062
DNMT1-209
ENST00000586800
583
25aa
ENSP00000465555
DNMT1-205
ENST00000586086
489
No protein
-
Pathway ID
Pathway Name
Source
ensgID
Trait
pValue
Pubmed ID
50694 Child Development Disorders, Pervasive 4.2719400E-005 - 70591 Hydrochlorothiazide 3E-6 23400010 70592 Triglycerides 3E-6 23400010
ensgID SNP Chromosome Position
Trait PubmedID Or or BEAT
EFO ID
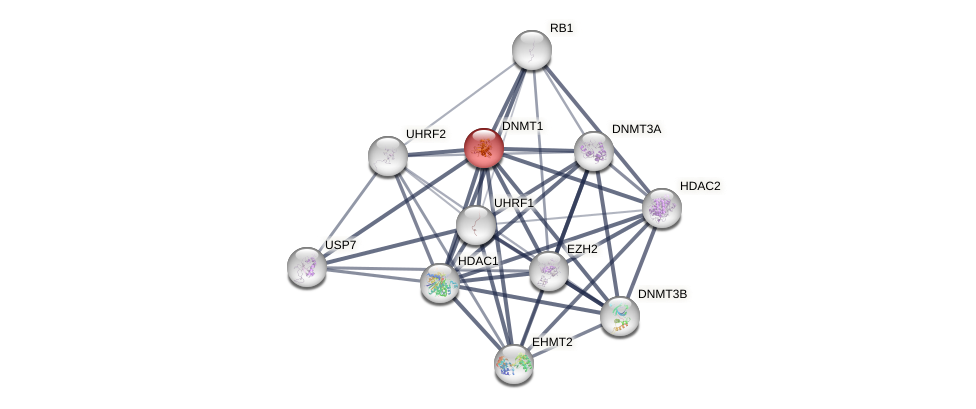
Protein-Protein Interaction (PPI)
Ensembl ID
Source
Target
Species
Protein ID
Perc_pos
Perc_id
Species
Protein ID
Perc_pos
Perc_id
ENSG00000130816 homo_sapiens ENSP00000367030 47.8261 27.3657 homo_sapiens ENSP00000352516 11.4583 6.55637
Ensembl ID
Source
Target
Species
Protein ID
Perc_pos
Perc_id
Species
Protein ID
Perc_pos
Perc_id
ENSG00000130816 homo_sapiens ENSP00000352516 96.1397 95.2206 nomascus_leucogenys ENSNLEP00000027390 95.322 94.4107 ENSG00000130816 homo_sapiens ENSP00000352516 96.9976 96.8137 pan_paniscus ENSPPAP00000008604 96.7013 96.518 ENSG00000130816 homo_sapiens ENSP00000352516 99.7549 99.4485 pongo_abelii ENSPPYP00000010706 99.7549 99.4485 ENSG00000130816 homo_sapiens ENSP00000352516 99.4485 99.2647 pan_troglodytes ENSPTRP00000085699 96.9534 96.7742 ENSG00000130816 homo_sapiens ENSP00000352516 99.4485 99.2647 pan_troglodytes ENSPTRP00000066034 96.9534 96.7742 ENSG00000130816 homo_sapiens ENSP00000352516 96.5074 96.0784 gorilla_gorilla ENSGGOP00000016186 94.994 94.5718 ENSG00000130816 homo_sapiens ENSP00000352516 73.1618 64.0931 ornithorhynchus_anatinus ENSOANP00000019393 79.8662 69.9666 ENSG00000130816 homo_sapiens ENSP00000352516 79.8407 72.7328 sarcophilus_harrisii ENSSHAP00000010299 75.2309 68.5335 ENSG00000130816 homo_sapiens ENSP00000352516 25.4289 21.5074 sarcophilus_harrisii ENSSHAP00000029869 81.0547 68.5547 ENSG00000130816 homo_sapiens ENSP00000352516 55.0858 48.223 notamacropus_eugenii ENSMEUP00000007371 55.8038 48.8516 ENSG00000130816 homo_sapiens ENSP00000352516 35.9681 32.9044 notamacropus_eugenii ENSMEUP00000011543 57.3242 52.4414 ENSG00000130816 homo_sapiens ENSP00000352516 57.8431 54.7181 choloepus_hoffmanni ENSCHOP00000008420 64.2177 60.7483 ENSG00000130816 homo_sapiens ENSP00000352516 80.7598 75.8578 dasypus_novemcinctus ENSDNOP00000002571 84.0025 78.9038 ENSG00000130816 homo_sapiens ENSP00000352516 51.1642 44.9755 erinaceus_europaeus ENSEEUP00000010530 66.7466 58.6731 ENSG00000130816 homo_sapiens ENSP00000352516 60.4167 54.8407 echinops_telfairi ENSETEP00000010201 62.4051 56.6456 ENSG00000130816 homo_sapiens ENSP00000352516 97.3652 96.0172 callithrix_jacchus ENSCJAP00000010808 98.3292 96.9678 ENSG00000130816 homo_sapiens ENSP00000352516 98.652 98.223 cercocebus_atys ENSCATP00000013989 98.652 98.223 ENSG00000130816 homo_sapiens ENSP00000352516 98.0392 97.6716 macaca_fascicularis ENSMFAP00000013070 93.1858 92.8363 ENSG00000130816 homo_sapiens ENSP00000352516 98.0392 97.6716 macaca_mulatta ENSMMUP00000019170 93.1858 92.8363 ENSG00000130816 homo_sapiens ENSP00000352516 97.549 97.1201 macaca_nemestrina ENSMNEP00000009892 92.9907 92.5818 ENSG00000130816 homo_sapiens ENSP00000352516 97.9167 97.6716 papio_anubis ENSPANP00000000133 98.8861 98.6386 ENSG00000130816 homo_sapiens ENSP00000352516 94.9755 94.1789 mandrillus_leucophaeus ENSMLEP00000029369 97.4843 96.6667 ENSG00000130816 homo_sapiens ENSP00000352516 84.4363 79.7181 tursiops_truncatus ENSTTRP00000002979 85.4839 80.7072 ENSG00000130816 homo_sapiens ENSP00000352516 87.5 84.3137 vulpes_vulpes ENSVVUP00000040627 94.8207 91.3679 ENSG00000130816 homo_sapiens ENSP00000352516 86.6422 77.3284 mus_spretus MGP_SPRETEiJ_P0090468 87.284 77.9012 ENSG00000130816 homo_sapiens ENSP00000352516 94.6078 90.1348 equus_asinus ENSEASP00005015119 90.2396 85.9731 ENSG00000130816 homo_sapiens ENSP00000352516 91.1152 86.2745 capra_hircus ENSCHIP00000027359 93.4632 88.4978 ENSG00000130816 homo_sapiens ENSP00000352516 94.1789 91.1765 loxodonta_africana ENSLAFP00000002359 94.4103 91.4005 ENSG00000130816 homo_sapiens ENSP00000352516 94.6691 90.9314 panthera_leo ENSPLOP00000004442 90.3509 86.7836 ENSG00000130816 homo_sapiens ENSP00000352516 94.6078 90.4412 ailuropoda_melanoleuca ENSAMEP00000012576 92.9001 88.8087 ENSG00000130816 homo_sapiens ENSP00000352516 93.1985 88.3578 bos_indicus_hybrid ENSBIXP00005042727 89.0515 84.4262 ENSG00000130816 homo_sapiens ENSP00000352516 94.5466 90.0123 equus_caballus ENSECAP00000035267 90.1812 85.8562 ENSG00000130816 homo_sapiens ENSP00000352516 94.3627 90.2574 canis_lupus_familiaris ENSCAFP00845025238 95.5928 91.4339 ENSG00000130816 homo_sapiens ENSP00000352516 94.6691 90.9314 panthera_pardus ENSPPRP00000008863 95.7843 92.0025 ENSG00000130816 homo_sapiens ENSP00000352516 85.5392 81.4338 marmota_marmota_marmota ENSMMMP00000019143 92.8191 88.3644 ENSG00000130816 homo_sapiens ENSP00000352516 93.5049 88.5417 balaenoptera_musculus ENSBMSP00010016360 94.1975 89.1975 ENSG00000130816 homo_sapiens ENSP00000352516 86.2132 79.5343 cavia_porcellus ENSCPOP00000019013 87.9375 81.125 ENSG00000130816 homo_sapiens ENSP00000352516 94.6691 90.9314 felis_catus ENSFCAP00000044002 90.3509 86.7836 ENSG00000130816 homo_sapiens ENSP00000352516 56.0662 54.2892 ursus_americanus ENSUAMP00000008485 93.2722 90.316 ENSG00000130816 homo_sapiens ENSP00000352516 75 67.0343 monodelphis_domestica ENSMODP00000053416 84.4138 75.4483 ENSG00000130816 homo_sapiens ENSP00000352516 80.6373 72.7941 monodelphis_domestica ENSMODP00000006657 86.9221 78.4676 ENSG00000130816 homo_sapiens ENSP00000352516 93.076 88.2966 bison_bison_bison ENSBBBP00000007175 94.2308 89.3921 ENSG00000130816 homo_sapiens ENSP00000352516 93.3824 88.4804 monodon_monoceros ENSMMNP00015029043 94.4238 89.4672 ENSG00000130816 homo_sapiens ENSP00000352516 93.1985 88.174 sus_scrofa ENSSSCP00000014522 90.6436 85.7569 ENSG00000130816 homo_sapiens ENSP00000352516 95.5882 92.5858 microcebus_murinus ENSMICP00000009338 96.7142 93.6764 ENSG00000130816 homo_sapiens ENSP00000352516 87.1936 80.8824 octodon_degus ENSODEP00000023082 90.3492 83.8095 ENSG00000130816 homo_sapiens ENSP00000352516 87.4387 81.0049 jaculus_jaculus ENSJJAP00000020454 92.0052 85.2353 ENSG00000130816 homo_sapiens ENSP00000352516 93.2598 88.2966 ovis_aries_rambouillet ENSOARP00020004700 89.1101 84.3677 ENSG00000130816 homo_sapiens ENSP00000352516 87.6226 83.6397 ursus_maritimus ENSUMAP00000034894 95.7803 91.4267 ENSG00000130816 homo_sapiens ENSP00000352516 79.0441 73.5294 ochotona_princeps ENSOPRP00000006465 80.0248 74.4417 ENSG00000130816 homo_sapiens ENSP00000352516 93.3824 88.5417 delphinapterus_leucas ENSDLEP00000011773 87.4856 82.9506 ENSG00000130816 homo_sapiens ENSP00000352516 93.2598 88.4804 physeter_catodon ENSPCTP00005009443 94.5342 89.6894 ENSG00000130816 homo_sapiens ENSP00000352516 80.6373 71.875 mus_caroli MGP_CAROLIEiJ_P0086915 87.6165 78.0959 ENSG00000130816 homo_sapiens ENSP00000352516 86.7647 77.5123 mus_musculus ENSMUSP00000004202 87.4074 78.0864 ENSG00000130816 homo_sapiens ENSP00000352516 84.6814 78.5539 dipodomys_ordii ENSDORP00000013274 87.8576 81.5003 ENSG00000130816 homo_sapiens ENSP00000352516 92.7083 88.6642 mustela_putorius_furo ENSMPUP00000006775 90.3823 86.4397 ENSG00000130816 homo_sapiens ENSP00000352516 97.4877 96.0172 aotus_nancymaae ENSANAP00000013919 98.2705 96.7881 ENSG00000130816 homo_sapiens ENSP00000352516 97.0588 95.7721 saimiri_boliviensis_boliviensis ENSSBOP00000038451 92.4154 91.1902 ENSG00000130816 homo_sapiens ENSP00000352516 48.3456 45.6495 vicugna_pacos ENSVPAP00000005678 50.6418 47.8177 ENSG00000130816 homo_sapiens ENSP00000352516 98.1005 97.549 chlorocebus_sabaeus ENSCSAP00000003647 99.0718 98.5149 ENSG00000130816 homo_sapiens ENSP00000352516 69.1176 66.1152 tupaia_belangeri ENSTBEP00000006380 69.5009 66.4818 ENSG00000130816 homo_sapiens ENSP00000352516 85.1103 77.3284 mesocricetus_auratus ENSMAUP00000022081 86.8668 78.9243 ENSG00000130816 homo_sapiens ENSP00000352516 93.3824 88.4804 phocoena_sinus ENSPSNP00000027437 94.016 89.0808 ENSG00000130816 homo_sapiens ENSP00000352516 94.0564 89.5833 camelus_dromedarius ENSCDRP00005009069 95.1053 90.5824 ENSG00000130816 homo_sapiens ENSP00000352516 82.4755 80.8824 carlito_syrichta ENSTSYP00000008234 97.8909 96 ENSG00000130816 homo_sapiens ENSP00000352516 88.7868 84.375 panthera_tigris_altaica ENSPTIP00000017059 89.2241 84.7906 ENSG00000130816 homo_sapiens ENSP00000352516 87.0711 78.4314 rattus_norvegicus ENSRNOP00000063831 87.6619 78.9636 ENSG00000130816 homo_sapiens ENSP00000352516 93.9951 90.8701 prolemur_simus ENSPSMP00000029456 93.0261 89.9333 ENSG00000130816 homo_sapiens ENSP00000352516 86.0294 77.8186 microtus_ochrogaster ENSMOCP00000014440 86.8275 78.5405 ENSG00000130816 homo_sapiens ENSP00000352516 74.9387 68.8726 sorex_araneus ENSSARP00000006203 78.0971 71.7752 ENSG00000130816 homo_sapiens ENSP00000352516 73.8971 63.4804 peromyscus_maniculatus_bairdii ENSPEMP00000031510 80.3464 69.0207 ENSG00000130816 homo_sapiens ENSP00000352516 87.0098 79.2892 peromyscus_maniculatus_bairdii ENSPEMP00000003713 88.0893 80.2729 ENSG00000130816 homo_sapiens ENSP00000352516 91.0539 85.9681 bos_mutus ENSBMUP00000027880 87.6179 82.7241 ENSG00000130816 homo_sapiens ENSP00000352516 86.5809 77.3897 mus_spicilegus ENSMSIP00000020610 87.2222 77.963 ENSG00000130816 homo_sapiens ENSP00000352516 83.8848 75.3064 cricetulus_griseus_chok1gshd ENSCGRP00001024361 86.6456 77.7848 ENSG00000130816 homo_sapiens ENSP00000352516 92.5858 88.5417 pteropus_vampyrus ENSPVAP00000003477 93.8509 89.7516 ENSG00000130816 homo_sapiens ENSP00000352516 89.277 82.7206 myotis_lucifugus ENSMLUP00000001254 91.4053 84.6926 ENSG00000130816 homo_sapiens ENSP00000352516 75.6127 68.6274 vombatus_ursinus ENSVURP00010032699 89.2263 80.9834 ENSG00000130816 homo_sapiens ENSP00000352516 27.5123 23.7745 vombatus_ursinus ENSVURP00010032703 80.466 69.534 ENSG00000130816 homo_sapiens ENSP00000352516 94.7917 90.5024 neovison_vison ENSNVIP00000030865 95.8488 91.5118 ENSG00000130816 homo_sapiens ENSP00000352516 93.1985 88.3578 bos_taurus ENSBTAP00000003549 92.5182 87.7129 ENSG00000130816 homo_sapiens ENSP00000352516 73.1618 68.75 heterocephalus_glaber_female ENSHGLP00000046148 92.4149 86.8421 ENSG00000130816 homo_sapiens ENSP00000352516 85.9681 85.5392 rhinopithecus_bieti ENSRBIP00000028369 94.6694 94.197 ENSG00000130816 homo_sapiens ENSP00000352516 66.3603 60.7843 canis_lupus_dingo ENSCAFP00020031226 80.9417 74.1405 ENSG00000130816 homo_sapiens ENSP00000352516 93.3211 88.7255 sciurus_vulgaris ENSSVLP00005025163 94.5376 89.8821 ENSG00000130816 homo_sapiens ENSP00000352516 80.7598 72.7941 phascolarctos_cinereus ENSPCIP00000024931 87.0542 78.4676 ENSG00000130816 homo_sapiens ENSP00000352516 26.5319 23.4681 phascolarctos_cinereus ENSPCIP00000024933 82.6336 73.0916 ENSG00000130816 homo_sapiens ENSP00000352516 91.7279 89.0931 procavia_capensis ENSPCAP00000010493 92.1798 89.532 ENSG00000130816 homo_sapiens ENSP00000352516 85.7843 79.3505 nannospalax_galili ENSNGAP00000007053 87.995 81.3953 ENSG00000130816 homo_sapiens ENSP00000352516 93.1985 88.3578 bos_grunniens ENSBGRP00000003888 94.4134 89.5096 ENSG00000130816 homo_sapiens ENSP00000352516 91.2377 84.6814 chinchilla_lanigera ENSCLAP00000013845 90.6269 84.1144 ENSG00000130816 homo_sapiens ENSP00000352516 92.7696 87.8676 cervus_hanglu_yarkandensis ENSCHYP00000002297 93.9206 88.9578 ENSG00000130816 homo_sapiens ENSP00000352516 93.076 88.3578 catagonus_wagneri ENSCWAP00000005962 92.3966 87.7129 ENSG00000130816 homo_sapiens ENSP00000352516 86.8873 82.1078 ictidomys_tridecemlineatus ENSSTOP00000025333 92.5587 87.4674 ENSG00000130816 homo_sapiens ENSP00000352516 93.076 89.7672 otolemur_garnettii ENSOGAP00000001132 95.7152 92.3125 ENSG00000130816 homo_sapiens ENSP00000352516 97.3652 95.8333 cebus_imitator ENSCCAP00000033707 97.6044 96.0688 ENSG00000130816 homo_sapiens ENSP00000352516 27.0221 25.6127 cebus_imitator ENSCCAP00000037775 85.1351 80.695 ENSG00000130816 homo_sapiens ENSP00000352516 79.2279 74.5098 urocitellus_parryii ENSUPAP00010000999 91.2491 85.8151 ENSG00000130816 homo_sapiens ENSP00000352516 91.4216 86.3358 moschus_moschiferus ENSMMSP00000003711 90.644 85.6015 ENSG00000130816 homo_sapiens ENSP00000352516 41.1765 40.7475 rhinopithecus_roxellana ENSRROP00000042387 97.2504 96.2373 ENSG00000130816 homo_sapiens ENSP00000352516 98.0392 97.6103 rhinopithecus_roxellana ENSRROP00000029954 96.0961 95.6757 ENSG00000130816 homo_sapiens ENSP00000352516 87.2549 77.8799 mus_pahari MGP_PahariEiJ_P0022563 87.9555 78.5052 ENSG00000130816 homo_sapiens ENSP00000352516 96.3848 93.5049 propithecus_coquereli ENSPCOP00000013513 97.3994 94.4892 ENSG00000130816 homo_sapiens ENSP00000352516 55.0858 44.424 ciona_intestinalis ENSCINP00000010797 71.5195 57.677 ENSG00000130816 homo_sapiens ENSP00000352516 27.7574 25.7966 petromyzon_marinus ENSPMAP00000002859 86.4504 80.3435 ENSG00000130816 homo_sapiens ENSP00000352516 6.37255 5.14706 eptatretus_burgeri ENSEBUP00000010816 68.4211 55.2632 ENSG00000130816 homo_sapiens ENSP00000352516 46.3848 38.6029 eptatretus_burgeri ENSEBUP00000027356 72.0266 59.9429 ENSG00000130816 homo_sapiens ENSP00000352516 16.299 14.4608 eptatretus_burgeri ENSEBUP00000003812 79.403 70.4478 ENSG00000130816 homo_sapiens ENSP00000352516 75.8578 65.7476 callorhinchus_milii ENSCMIP00000017481 82.0411 71.1067 ENSG00000130816 homo_sapiens ENSP00000352516 65.5024 58.2721 latimeria_chalumnae ENSLACP00000012502 83.9749 74.7054 ENSG00000130816 homo_sapiens ENSP00000352516 75.4289 65.5637 lepisosteus_oculatus ENSLOCP00000009053 77.8621 67.6787 ENSG00000130816 homo_sapiens ENSP00000352516 74.2647 63.4191 clupea_harengus ENSCHAP00000035180 79.9472 68.2718 ENSG00000130816 homo_sapiens ENSP00000352516 74.8774 65.4412 danio_rerio ENSDARP00000013243 81.4667 71.2 ENSG00000130816 homo_sapiens ENSP00000352516 75 65.7476 carassius_auratus ENSCARP00000091344 81.5456 71.4857 ENSG00000130816 homo_sapiens ENSP00000352516 74.8774 65.6863 carassius_auratus ENSCARP00000088046 81.1421 71.1819 ENSG00000130816 homo_sapiens ENSP00000352516 74.6936 65.2574 carassius_auratus ENSCARP00000121032 80.8355 70.6233 ENSG00000130816 homo_sapiens ENSP00000352516 73.7132 64.2157 astyanax_mexicanus ENSAMXP00000012561 81.3938 70.9066 ENSG00000130816 homo_sapiens ENSP00000352516 75.0613 65.5637 ictalurus_punctatus ENSIPUP00000007009 81.3413 71.0491 ENSG00000130816 homo_sapiens ENSP00000352516 75.0613 64.4608 esox_lucius ENSELUP00000024960 80.0654 68.7582 ENSG00000130816 homo_sapiens ENSP00000352516 73.1618 63.7255 electrophorus_electricus ENSEEEP00000041978 80.188 69.8455 ENSG00000130816 homo_sapiens ENSP00000352516 74.0809 62.8064 oncorhynchus_kisutch ENSOKIP00005004743 78.7109 66.7318 ENSG00000130816 homo_sapiens ENSP00000352516 74.8774 63.7868 oncorhynchus_kisutch ENSOKIP00005082035 79.8693 68.0392 ENSG00000130816 homo_sapiens ENSP00000352516 75.4289 63.9706 oncorhynchus_mykiss ENSOMYP00000092105 80.6684 68.4142 ENSG00000130816 homo_sapiens ENSP00000352516 74.0196 63.3578 oncorhynchus_mykiss ENSOMYP00000092658 78.9542 67.5817 ENSG00000130816 homo_sapiens ENSP00000352516 74.6936 63.2966 salmo_salar ENSSSAP00000020322 80.6217 68.3201 ENSG00000130816 homo_sapiens ENSP00000352516 75.1226 64.1544 salmo_salar ENSSSAP00000038496 80.0784 68.3867 ENSG00000130816 homo_sapiens ENSP00000352516 75.3064 63.848 salmo_trutta ENSSTUP00000032224 80.4319 68.1937 ENSG00000130816 homo_sapiens ENSP00000352516 74.8162 64.0319 salmo_trutta ENSSTUP00000040089 76.6478 65.5995 ENSG00000130816 homo_sapiens ENSP00000352516 73.8971 62.3775 gadus_morhua ENSGMOP00000035599 77.3573 65.2983 ENSG00000130816 homo_sapiens ENSP00000352516 74.6324 63.6642 fundulus_heteroclitus ENSFHEP00000000830 80.984 69.0824 ENSG00000130816 homo_sapiens ENSP00000352516 74.4485 63.6029 poecilia_reticulata ENSPREP00000009697 80.5703 68.8329 ENSG00000130816 homo_sapiens ENSP00000352516 74.2034 63.6029 xiphophorus_maculatus ENSXMAP00000040629 80.7872 69.2462 ENSG00000130816 homo_sapiens ENSP00000352516 74.5711 63.7868 oryzias_latipes ENSORLP00000034016 81.3503 69.5856 ENSG00000130816 homo_sapiens ENSP00000352516 74.5711 62.8064 cyclopterus_lumpus ENSCLMP00005047527 81.1333 68.3333 ENSG00000130816 homo_sapiens ENSP00000352516 74.8162 63.9706 oreochromis_niloticus ENSONIP00000029113 80.8609 69.1391 ENSG00000130816 homo_sapiens ENSP00000352516 74.5711 63.7868 haplochromis_burtoni ENSHBUP00000034119 80.7565 69.0776 ENSG00000130816 homo_sapiens ENSP00000352516 74.5098 63.7255 astatotilapia_calliptera ENSACLP00000027974 80.6901 69.0113 ENSG00000130816 homo_sapiens ENSP00000352516 74.8774 64.0931 sparus_aurata ENSSAUP00010043284 79.4538 68.0104 ENSG00000130816 homo_sapiens ENSP00000352516 75.2451 64.7672 lates_calcarifer ENSLCAP00010056006 81.2707 69.9537 ENSG00000130816 homo_sapiens ENSP00000352516 78.5539 68.75 xenopus_tropicalis ENSXETP00000047703 85.9249 75.2011 ENSG00000130816 homo_sapiens ENSP00000352516 74.8162 67.1569 chrysemys_picta_bellii ENSCPBP00000006854 87.8417 78.8489 ENSG00000130816 homo_sapiens ENSP00000352516 64.4608 56.1887 sphenodon_punctatus ENSSPUP00000013434 79.0977 68.9474 ENSG00000130816 homo_sapiens ENSP00000352516 74.3873 65.6863 laticauda_laticaudata ENSLLTP00000000076 86.6524 76.5168 ENSG00000130816 homo_sapiens ENSP00000352516 76.5931 67.7696 notechis_scutatus ENSNSUP00000019679 85.2079 75.392 ENSG00000130816 homo_sapiens ENSP00000352516 79.6569 69.7304 pseudonaja_textilis ENSPTXP00000006608 85.3578 74.7209 ENSG00000130816 homo_sapiens ENSP00000352516 73.223 65.8088 anas_platyrhynchos_platyrhynchos ENSAPLP00000002034 85.5404 76.879 ENSG00000130816 homo_sapiens ENSP00000352516 79.1054 70.5882 gallus_gallus ENSGALP00010004045 80.738 72.045 ENSG00000130816 homo_sapiens ENSP00000352516 62.2549 54.7794 meleagris_gallopavo ENSMGAP00000029840 80.1262 70.5047 ENSG00000130816 homo_sapiens ENSP00000352516 56.1275 48.9583 serinus_canaria ENSSCAP00000010650 80.2102 69.965 ENSG00000130816 homo_sapiens ENSP00000352516 79.1054 69.424 anolis_carolinensis ENSACAP00000004041 82.3342 72.2577 ENSG00000130816 homo_sapiens ENSP00000352516 74.326 63.7255 neolamprologus_brichardi ENSNBRP00000012739 81.0287 69.4723 ENSG00000130816 homo_sapiens ENSP00000352516 79.7181 69.6691 naja_naja ENSNNAP00000029453 85.4235 74.6553 ENSG00000130816 homo_sapiens ENSP00000352516 79.8407 69.9755 podarcis_muralis ENSPMRP00000036840 85.4987 74.9344 ENSG00000130816 homo_sapiens ENSP00000352516 72.3652 63.5417 geospiza_fortis ENSGFOP00000006527 80.5593 70.7367 ENSG00000130816 homo_sapiens ENSP00000352516 75.1838 66.8505 taeniopygia_guttata ENSTGUP00000031616 81.6367 72.5882 ENSG00000130816 homo_sapiens ENSP00000352516 72.2426 63.7255 erpetoichthys_calabaricus ENSECRP00000024237 83.8549 73.9687 ENSG00000130816 homo_sapiens ENSP00000352516 74.8774 63.9706 kryptolebias_marmoratus ENSKMAP00000023146 81.0883 69.2767 ENSG00000130816 homo_sapiens ENSP00000352516 78.8603 69.3015 salvator_merianae ENSSMRP00000021541 84.6154 74.359 ENSG00000130816 homo_sapiens ENSP00000352516 75.1838 63.848 amphilophus_citrinellus ENSACIP00000010933 81.42 69.144 ENSG00000130816 homo_sapiens ENSP00000352516 74.326 63.5417 dicentrarchus_labrax ENSDLAP00005073637 79.8026 68.2237 ENSG00000130816 homo_sapiens ENSP00000352516 74.3873 63.7255 amphiprion_percula ENSAPEP00000014908 80.3974 68.8742 ENSG00000130816 homo_sapiens ENSP00000352516 69.3015 60.4167 oryzias_sinensis ENSOSIP00000013275 82.9787 72.3404 ENSG00000130816 homo_sapiens ENSP00000352516 64.7059 55.8824 pelodiscus_sinensis ENSPSIP00000003025 76.2455 65.8484 ENSG00000130816 homo_sapiens ENSP00000352516 79.1054 70.6495 gopherus_evgoodei ENSGEVP00005027808 84.7113 75.6562 ENSG00000130816 homo_sapiens ENSP00000352516 78.0024 69.9142 chelonoidis_abingdonii ENSCABP00000024081 84.0819 75.3633 ENSG00000130816 homo_sapiens ENSP00000352516 75.0613 64.3382 seriola_dumerili ENSSDUP00000029478 81.3413 69.7211 ENSG00000130816 homo_sapiens ENSP00000352516 74.9387 63.0515 cottoperca_gobio ENSCGOP00000018946 81.3706 68.4631 ENSG00000130816 homo_sapiens ENSP00000352516 57.7206 51.7157 struthio_camelus_australis ENSSCUP00000020652 85.0181 76.1733 ENSG00000130816 homo_sapiens ENSP00000352516 75.3064 67.7696 anser_brachyrhynchus ENSABRP00000001040 88.1004 79.2832 ENSG00000130816 homo_sapiens ENSP00000352516 74.7549 63.2353 gasterosteus_aculeatus ENSGACP00000018172 81.0093 68.5259 ENSG00000130816 homo_sapiens ENSP00000352516 69.9142 59.8652 cynoglossus_semilaevis ENSCSEP00000032828 79.5676 68.1311 ENSG00000130816 homo_sapiens ENSP00000352516 74.7549 63.9706 poecilia_formosa ENSPFOP00000006222 81.171 69.4611 ENSG00000130816 homo_sapiens ENSP00000352516 74.7549 64.3995 myripristis_murdjan ENSMMDP00005001728 80.9019 69.695 ENSG00000130816 homo_sapiens ENSP00000352516 75.1226 63.848 sander_lucioperca ENSSLUP00000021297 81.6789 69.4204 ENSG00000130816 homo_sapiens ENSP00000352516 76.4093 68.3824 strigops_habroptila ENSSHBP00005019369 76.3625 68.3405 ENSG00000130816 homo_sapiens ENSP00000352516 74.5098 63.848 amphiprion_ocellaris ENSAOCP00000020885 80.5298 69.0066 ENSG00000130816 homo_sapiens ENSP00000352516 79.7794 70.8946 terrapene_carolina_triunguis ENSTMTP00000004785 85.8839 76.3193 ENSG00000130816 homo_sapiens ENSP00000352516 75.3676 64.0931 hucho_hucho ENSHHUP00000073641 81.0277 68.9065 ENSG00000130816 homo_sapiens ENSP00000352516 71.5074 60.6618 hucho_hucho ENSHHUP00000070445 79.0115 67.0278 ENSG00000130816 homo_sapiens ENSP00000352516 75.3676 64.3995 betta_splendens ENSBSLP00000033092 81.457 69.6026 ENSG00000130816 homo_sapiens ENSP00000352516 75.2451 65.8701 pygocentrus_nattereri ENSPNAP00000030448 81.2707 71.1449 ENSG00000130816 homo_sapiens ENSP00000352516 75.3064 64.8284 anabas_testudineus ENSATEP00000030651 81.4987 70.1591 ENSG00000130816 homo_sapiens ENSP00000352516 54.2279 43.8113 ciona_savignyi ENSCSAVP00000010172 70.1824 56.701 ENSG00000130816 homo_sapiens ENSP00000352516 74.6936 64.1544 seriola_lalandi_dorsalis ENSSLDP00000020457 81.1585 69.7071 ENSG00000130816 homo_sapiens ENSP00000352516 71.6299 61.2745 labrus_bergylta ENSLBEP00000027458 82.5565 70.6215 ENSG00000130816 homo_sapiens ENSP00000352516 74.6324 63.848 pundamilia_nyererei ENSPNYP00000018171 81.1459 69.4204 ENSG00000130816 homo_sapiens ENSP00000352516 75.4902 64.4608 mastacembelus_armatus ENSMAMP00000018326 81.8061 69.8539 ENSG00000130816 homo_sapiens ENSP00000352516 74.7549 63.7868 stegastes_partitus ENSSPAP00000000592 81.117 69.2154 ENSG00000130816 homo_sapiens ENSP00000352516 70.7721 60.2941 oncorhynchus_tshawytscha ENSOTSP00005073573 79.8755 68.0498 ENSG00000130816 homo_sapiens ENSP00000352516 75.2451 63.9093 oncorhynchus_tshawytscha ENSOTSP00005054272 80.1567 68.0809 ENSG00000130816 homo_sapiens ENSP00000352516 74.5711 63.9093 acanthochromis_polyacanthus ENSAPOP00000014617 80.6494 69.1186 ENSG00000130816 homo_sapiens ENSP00000352516 74.7549 67.2181 aquila_chrysaetos_chrysaetos ENSACCP00020001916 81.8792 73.6242 ENSG00000130816 homo_sapiens ENSP00000352516 74.326 63.9706 cyprinodon_variegatus ENSCVAP00000027344 80.8128 69.5536 ENSG00000130816 homo_sapiens ENSP00000352516 70.5882 61.7647 denticeps_clupeoides ENSDCDP00000026007 76.3926 66.8435 ENSG00000130816 homo_sapiens ENSP00000352516 74.6936 63.7868 larimichthys_crocea ENSLCRP00005013625 80.6217 68.8492 ENSG00000130816 homo_sapiens ENSP00000352516 74.0196 62.9902 takifugu_rubripes ENSTRUP00000020322 77.5353 65.982 ENSG00000130816 homo_sapiens ENSP00000352516 62.7451 56.4951 ficedula_albicollis ENSFALP00000008124 84.4188 76.0099 ENSG00000130816 homo_sapiens ENSP00000352516 73.9583 63.3578 hippocampus_comes ENSHCOP00000013439 81.2795 69.6296 ENSG00000130816 homo_sapiens ENSP00000352516 74.9387 65.3799 cyprinus_carpio_carpio ENSCCRP00000176248 81.1008 70.756 ENSG00000130816 homo_sapiens ENSP00000352516 75.1226 65.9314 cyprinus_carpio_carpio ENSCCRP00000144653 79.7139 69.961 ENSG00000130816 homo_sapiens ENSP00000352516 62.9902 55.2696 coturnix_japonica ENSCJPP00005005178 80.1872 70.3588 ENSG00000130816 homo_sapiens ENSP00000352516 74.7549 63.9706 poecilia_latipinna ENSPLAP00000017030 81.171 69.4611 ENSG00000130816 homo_sapiens ENSP00000352516 72.3039 63.4191 sinocyclocheilus_grahami ENSSGRP00000090638 79.7836 69.9797 ENSG00000130816 homo_sapiens ENSP00000352516 74.5711 64.8897 sinocyclocheilus_grahami ENSSGRP00000037425 80.9714 70.4591 ENSG00000130816 homo_sapiens ENSP00000352516 74.6324 63.848 maylandia_zebra ENSMZEP00005025334 81.0919 69.3742 ENSG00000130816 homo_sapiens ENSP00000352516 68.3824 60.049 tetraodon_nigroviridis ENSTNIP00000019029 83.0357 72.9167 ENSG00000130816 homo_sapiens ENSP00000352516 26.1642 23.223 paramormyrops_kingsleyae ENSPKIP00000028718 85.9155 76.2575 ENSG00000130816 homo_sapiens ENSP00000352516 11.0907 10.1103 paramormyrops_kingsleyae ENSPKIP00000037174 89.1626 81.2808 ENSG00000130816 homo_sapiens ENSP00000352516 75.1838 65.3186 scleropages_formosus ENSSFOP00015048329 81.0436 70.4095 ENSG00000130816 homo_sapiens ENSP00000352516 75.674 66.6054 leptobrachium_leishanense ENSLLEP00000006783 82.6087 72.709 ENSG00000130816 homo_sapiens ENSP00000352516 75.0613 64.4608 scophthalmus_maximus ENSSMAP00000006735 81.4495 69.9468 ENSG00000130816 homo_sapiens ENSP00000352516 74.2647 63.2966 oryzias_melastigma ENSOMEP00000000064 80.9619 69.0047
Go ID
Go term
No. evidence
Entries
Species
Category
GO:0000122 involved_in negative regulation of transcription by RNA polymerase II 1 IEA Homo_sapiens(9606) Process GO:0001674 located_in female germ cell nucleus 1 IEA Homo_sapiens(9606) Component GO:0003677 enables DNA binding 2 IBA,IDA Homo_sapiens(9606) Function GO:0003723 enables RNA binding 1 IEA Homo_sapiens(9606) Function GO:0003886 enables DNA (cytosine-5-)-methyltransferase activity 2 IBA,IDA Homo_sapiens(9606) Function GO:0005515 enables protein binding 1 IPI Homo_sapiens(9606) Function GO:0005634 located_in nucleus 2 HDA,IBA Homo_sapiens(9606) Component GO:0005654 located_in nucleoplasm 1 TAS Homo_sapiens(9606) Component GO:0005657 located_in replication fork 1 IEA Homo_sapiens(9606) Component GO:0005721 part_of pericentric heterochromatin 1 IEA Homo_sapiens(9606) Component GO:0006346 involved_in DNA methylation-dependent heterochromatin formation 1 IEA Homo_sapiens(9606) Process GO:0006351 involved_in DNA-templated transcription 1 IEA Homo_sapiens(9606) Process GO:0008270 enables zinc ion binding 1 IEA Homo_sapiens(9606) Function GO:0008327 enables methyl-CpG binding 1 IEA Homo_sapiens(9606) Function GO:0009008 enables DNA-methyltransferase activity 1 IDA Homo_sapiens(9606) Function GO:0010424 involved_in DNA methylation on cytosine within a CG sequence 1 IBA Homo_sapiens(9606) Process GO:0010628 involved_in positive regulation of gene expression 1 IMP Homo_sapiens(9606) Process GO:0010629 involved_in negative regulation of gene expression 1 IMP Homo_sapiens(9606) Process GO:0043045 involved_in post-fertilization epigenetic regulation of gene expression 1 IEA Homo_sapiens(9606) Process GO:0044027 involved_in negative regulation of gene expression via CpG island methylation 2 IDA,IMP Homo_sapiens(9606) Process GO:0071230 involved_in cellular response to amino acid stimulus 1 IEA Homo_sapiens(9606) Process GO:0090116 involved_in C-5 methylation of cytosine 1 IEA Homo_sapiens(9606) Process GO:1903926 involved_in cellular response to bisphenol A 1 IEA Homo_sapiens(9606) Process GO:1904707 involved_in positive regulation of vascular associated smooth muscle cell proliferation 1 IMP Homo_sapiens(9606) Process GO:1905460 involved_in negative regulation of vascular associated smooth muscle cell apoptotic process 1 IMP Homo_sapiens(9606) Process GO:1905931 involved_in negative regulation of vascular associated smooth muscle cell differentiation involved in phenotypic switching 1 IMP Homo_sapiens(9606) Process GO:1990841 enables promoter-specific chromatin binding 1 IDA Homo_sapiens(9606) Function
Copyright © 2023, Bioinformatics Center, Sun Yat-sen Memorial Hospital, Sun Yat-sen University, China. All Rights Reserved.