RBP Type
N.A.
Diseases
Drug
Main interacting RNAs
Moonlighting functions
Localizations
BulkPerturb-seq
Ensembl ID YNL085W Gene ID
855639 Accession
-
Symbol
MKT1
Alias
NA
Full Name
NA
Status
Confidence
Length
2493 bases
Strand
Plus strand
Position
XIV : 467131 - 469623
RNA binding domain
MKT1_C , MKT1_N
Summary
Predicted to enable 5'-flap endonuclease activity. Involved in cellular response to DNA damage stimulus. Located in P-body and nuclear periphery. Part of polysome. [provided by Alliance of Genome Resources, Apr 2022]
RNA binding domains (RBDs)
Protein
Domain
Pfam ID
E-value
Domain number
YNL085W
MKT1_C
PF12246 9.7e-65
1
YNL085W
MKT1_N
PF12247 5.7e-30
2
RNA binding proteomes (RBPomes)
Pubmed ID
Full Name
Cell
Author
Time
Doi
Literatures on RNA binding capacity
Pubmed ID
Title
Author
Time
Journal
Name
Transcript ID
bp
Protein
Translation ID
-
YNL085W_mRNA
2493
830aa
YNL085W
Pathway ID
Pathway Name
Source
ensgID
Trait
pValue
Pubmed ID
ensgID SNP Chromosome Position
Trait PubmedID Or or BEAT
EFO ID
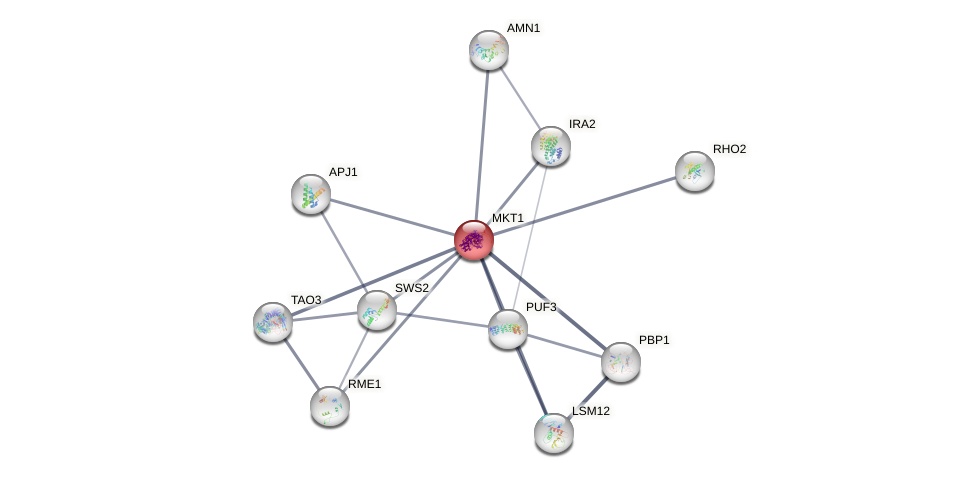
Protein-Protein Interaction (PPI)
Ensembl ID
Source
Target
Species
Protein ID
Perc_pos
Perc_id
Species
Protein ID
Perc_pos
Perc_id
YNL085W
Ensembl ID
Source
Target
Species
Protein ID
Perc_pos
Perc_id
Species
Protein ID
Perc_pos
Perc_id
YNL085W
Go ID
Go term
No. evidence
Entries
Species
Category
GO:0000932 located_in P-body 1 IDA Saccharomyces_cerevisiae(559292) Component GO:0003730 enables mRNA 3-UTR binding 1 IBA Saccharomyces_cerevisiae(559292) Function GO:0004518 enables nuclease activity 1 IEA Saccharomyces_cerevisiae(559292) Function GO:0004520 enables DNA endonuclease activity 1 ISS Saccharomyces_cerevisiae(559292) Function GO:0005634 is_active_in nucleus 1 IBA Saccharomyces_cerevisiae(559292) Component GO:0005737 located_in cytoplasm 2 HDA,IEA Saccharomyces_cerevisiae(559292) Component GO:0005829 located_in cytosol 1 IEA Saccharomyces_cerevisiae(559292) Component GO:0005844 part_of polysome 1 IDA Saccharomyces_cerevisiae(559292) Component GO:0006417 involved_in regulation of translation 1 IEA Saccharomyces_cerevisiae(559292) Process GO:0006974 involved_in DNA damage response 2 IEA,IMP Saccharomyces_cerevisiae(559292) Process GO:0010494 located_in cytoplasmic stress granule 1 HDA Saccharomyces_cerevisiae(559292) Component GO:0034399 located_in nuclear periphery 1 IDA Saccharomyces_cerevisiae(559292) Component
Copyright © 2023, Bioinformatics Center, Sun Yat-sen Memorial Hospital, Sun Yat-sen University, China. All Rights Reserved.