Welcome to
RBP World!
Description
| Ensembl ID | WBGene00000479 | Gene ID | 176061 | Accession | - |
| Symbol | cgh-1 | Alias | NA | Full Name | ATP-dependent RNA helicase cgh-1 |
| Status | Confidence | Length | 2513 bases | Strand | Minus strand |
| Position | III : 7495320 - 7497832 | RNA binding domain | DEAD | ||
| Summary | Predicted to enable RNA helicase activity and mRNA binding activity. Involved in several processes, including P-body assembly; determination of adult lifespan; and negative regulation of translation. Located in P granule; P-body; and cytoplasmic stress granule. Part of messenger ribonucleoprotein complex. Is expressed in several structures, including Z2; Z3; germ line; gonad; and neurons. Orthologous to human DDX6 (DEAD-box helicase 6). [provided by Alliance of Genome Resources, Apr 2022] | ||||
RNA binding domains (RBDs)
| Protein | Domain | Pfam ID | E-value | Domain number |
|---|---|---|---|---|
| C07H6 | DEAD | PF00270 | 4.6e-43 | 1 |
RNA binding proteomes (RBPomes)
| Pubmed ID | Full Name | Cell | Author | Time | Doi |
|---|
Literatures on RNA binding capacity
| Pubmed ID | Title | Author | Time | Journal |
|---|
Transcripts
| Name | Transcript ID | bp | Protein | Translation ID |
|---|---|---|---|---|
| cgh-1-201 | C07H6 | 2273 | 430aa | C07H6 |
Phenotypes
| ensgID | Trait | pValue | Pubmed ID |
|---|
GWAS
| ensgID | SNP | Chromosome | Position | Trait | PubmedID | Or or BEAT | EFO ID |
|---|
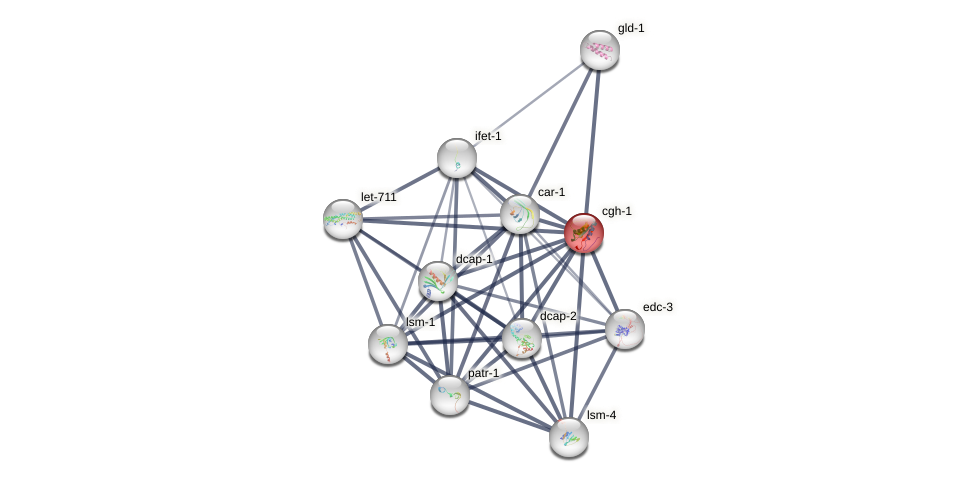
Protein-Protein Interaction (PPI)
Paralogs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| WBGene00000479 | caenorhabditis_elegans | C26D10.2a.1 | 43.2941 | 25.8824 | caenorhabditis_elegans | C07H6.5.1 | 42.7907 | 25.5814 |
| WBGene00000479 | caenorhabditis_elegans | C55B7.1.1 | 18.3778 | 9.95893 | caenorhabditis_elegans | C07H6.5.1 | 41.6279 | 22.5581 |
| WBGene00000479 | caenorhabditis_elegans | T21G5.3.1 | 23.46 | 12.713 | caenorhabditis_elegans | C07H6.5.1 | 41.6279 | 22.5581 |
| WBGene00000479 | caenorhabditis_elegans | B0414.6.1 | 25.1389 | 14.4444 | caenorhabditis_elegans | C07H6.5.1 | 42.093 | 24.186 |
| WBGene00000479 | caenorhabditis_elegans | Y38A10A.6.1 | 30.9615 | 15.7692 | caenorhabditis_elegans | C07H6.5.1 | 37.4419 | 19.0698 |
| WBGene00000479 | caenorhabditis_elegans | C14C11.6.1 | 30.6818 | 15.7197 | caenorhabditis_elegans | C07H6.5.1 | 37.6744 | 19.3023 |
| WBGene00000479 | caenorhabditis_elegans | F57B9.6a.1 | 50.2488 | 31.592 | caenorhabditis_elegans | C07H6.5.1 | 46.9767 | 29.5349 |
| WBGene00000479 | caenorhabditis_elegans | Y94H6A.5a.1 | 22.1818 | 12 | caenorhabditis_elegans | C07H6.5.1 | 42.5581 | 23.0233 |
| WBGene00000479 | caenorhabditis_elegans | ZK686.2.1 | 29.0051 | 14.8398 | caenorhabditis_elegans | C07H6.5.1 | 40 | 20.4651 |
| WBGene00000479 | caenorhabditis_elegans | Y71G12B.8.1 | 25.5751 | 15.1556 | caenorhabditis_elegans | C07H6.5.1 | 43.9535 | 26.0465 |
| WBGene00000479 | caenorhabditis_elegans | Y65B4A.6a.1 | 48.8722 | 29.5739 | caenorhabditis_elegans | C07H6.5.1 | 45.3488 | 27.4419 |
| WBGene00000479 | caenorhabditis_elegans | F33D11.10.1 | 48.8722 | 29.5739 | caenorhabditis_elegans | C07H6.5.1 | 45.3488 | 27.4419 |
| WBGene00000479 | caenorhabditis_elegans | ZK512.2a.1 | 26.8166 | 13.6678 | caenorhabditis_elegans | C07H6.5.1 | 36.0465 | 18.3721 |
| WBGene00000479 | caenorhabditis_elegans | R05D11.4.1 | 27.5387 | 16.0069 | caenorhabditis_elegans | C07H6.5.1 | 37.2093 | 21.6279 |
| WBGene00000479 | caenorhabditis_elegans | T26G10.1.1 | 38.2413 | 22.9039 | caenorhabditis_elegans | C07H6.5.1 | 43.4884 | 26.0465 |
| WBGene00000479 | caenorhabditis_elegans | Y55F3BR.1.1 | 22.2527 | 12.6374 | caenorhabditis_elegans | C07H6.5.1 | 37.6744 | 21.3953 |
| WBGene00000479 | caenorhabditis_elegans | Y54G11A.3.1 | 35.119 | 22.2222 | caenorhabditis_elegans | C07H6.5.1 | 41.1628 | 26.0465 |
| WBGene00000479 | caenorhabditis_elegans | Y23H5B.6a.1 | 25 | 14.3443 | caenorhabditis_elegans | C07H6.5.1 | 42.5581 | 24.4186 |
| WBGene00000479 | caenorhabditis_elegans | B0511.6.1 | 31.8015 | 18.3824 | caenorhabditis_elegans | C07H6.5.1 | 40.2326 | 23.2558 |
| WBGene00000479 | caenorhabditis_elegans | T07D4.4a.1 | 17.8082 | 10.4697 | caenorhabditis_elegans | C07H6.5.1 | 42.3256 | 24.8837 |
| WBGene00000479 | caenorhabditis_elegans | Y71H2AM.19.1 | 26.8362 | 15.3955 | caenorhabditis_elegans | C07H6.5.1 | 44.186 | 25.3488 |
| WBGene00000479 | caenorhabditis_elegans | F01F1.7.1 | 25.7534 | 13.8356 | caenorhabditis_elegans | C07H6.5.1 | 43.7209 | 23.4884 |
| WBGene00000479 | caenorhabditis_elegans | C24H12.4a.1 | 28.2334 | 16.4038 | caenorhabditis_elegans | C07H6.5.1 | 41.6279 | 24.186 |
| WBGene00000479 | caenorhabditis_elegans | F55F8.2a.1 | 21.9839 | 11.9303 | caenorhabditis_elegans | C07H6.5.1 | 38.1395 | 20.6977 |
| WBGene00000479 | caenorhabditis_elegans | H20J04.4b.1 | 30.8378 | 18.7166 | caenorhabditis_elegans | C07H6.5.1 | 40.2326 | 24.4186 |
| WBGene00000479 | caenorhabditis_elegans | H27M09.1.1 | 28.4127 | 16.9841 | caenorhabditis_elegans | C07H6.5.1 | 41.6279 | 24.8837 |
| WBGene00000479 | caenorhabditis_elegans | T06A10.1a.1 | 18.0884 | 11.3052 | caenorhabditis_elegans | C07H6.5.1 | 40.9302 | 25.5814 |
| WBGene00000479 | caenorhabditis_elegans | F53H1.1a.1 | 21.3402 | 11.0309 | caenorhabditis_elegans | C07H6.5.1 | 48.1395 | 24.8837 |
| WBGene00000479 | caenorhabditis_elegans | F58E10.3a.1 | 33.3333 | 19.9643 | caenorhabditis_elegans | C07H6.5.1 | 43.4884 | 26.0465 |
| WBGene00000479 | caenorhabditis_elegans | C46F11.4.1 | 21.9482 | 13.4402 | caenorhabditis_elegans | C07H6.5.1 | 41.3953 | 25.3488 |
Orthologs
| Ensembl ID | Source | Target | ||||||
|---|---|---|---|---|---|---|---|---|
| Species | Protein ID | Perc_pos | Perc_id | Species | Protein ID | Perc_pos | Perc_id | |
| WBGene00000479 | caenorhabditis_elegans | C07H6.5.1 | 81.1628 | 68.8372 | ciona_intestinalis | ENSCINP00000015240 | 80.2299 | 68.046 |
| WBGene00000479 | caenorhabditis_elegans | C07H6.5.1 | 85.3488 | 74.186 | drosophila_melanogaster | FBpp0079565 | 79.9564 | 69.4989 |
| WBGene00000479 | caenorhabditis_elegans | C07H6.5.1 | 76.0465 | 58.1395 | cervus_hanglu_yarkandensis | ENSCHYP00000018388 | 69.7228 | 53.3049 |
| WBGene00000479 | caenorhabditis_elegans | C07H6.5.1 | 77.907 | 63.9535 | saccharomyces_cerevisiae | YDL160C | 66.2055 | 54.3478 |
Gene Ontology
| Go ID | Go term | No. evidence | Entries | Species | Category |
|---|---|---|---|---|---|
| GO:0000166 | enables nucleotide binding | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0000932 | is_active_in P-body | 2 | IBA,IDA | Caenorhabditis_elegans(6239) | Component |
| GO:0003676 | enables nucleic acid binding | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0003723 | enables RNA binding | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0003724 | enables RNA helicase activity | 2 | IEA,ISS | Caenorhabditis_elegans(6239) | Function |
| GO:0003729 | enables mRNA binding | 1 | IBA | Caenorhabditis_elegans(6239) | Function |
| GO:0004386 | enables helicase activity | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0005515 | enables protein binding | 1 | IPI | Caenorhabditis_elegans(6239) | Function |
| GO:0005524 | enables ATP binding | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0005737 | located_in cytoplasm | 1 | IEA | Caenorhabditis_elegans(6239) | Component |
| GO:0006915 | involved_in apoptotic process | 1 | IEA | Caenorhabditis_elegans(6239) | Process |
| GO:0007276 | involved_in gamete generation | 1 | IMP | Caenorhabditis_elegans(6239) | Process |
| GO:0007283 | involved_in spermatogenesis | 1 | IEA | Caenorhabditis_elegans(6239) | Process |
| GO:0008340 | involved_in determination of adult lifespan | 1 | IMP | Caenorhabditis_elegans(6239) | Process |
| GO:0010494 | is_active_in cytoplasmic stress granule | 2 | IBA,IDA | Caenorhabditis_elegans(6239) | Component |
| GO:0016071 | involved_in mRNA metabolic process | 2 | IMP,TAS | Caenorhabditis_elegans(6239) | Process |
| GO:0016787 | enables hydrolase activity | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0016887 | enables ATP hydrolysis activity | 1 | IEA | Caenorhabditis_elegans(6239) | Function |
| GO:0017148 | involved_in negative regulation of translation | 2 | IBA,IMP | Caenorhabditis_elegans(6239) | Process |
| GO:0030154 | involved_in cell differentiation | 1 | IEA | Caenorhabditis_elegans(6239) | Process |
| GO:0033962 | involved_in P-body assembly | 2 | IBA,IMP | Caenorhabditis_elegans(6239) | Process |
| GO:0034063 | involved_in stress granule assembly | 1 | IBA | Caenorhabditis_elegans(6239) | Process |
| GO:0043066 | involved_in negative regulation of apoptotic process | 1 | IMP | Caenorhabditis_elegans(6239) | Process |
| GO:0043186 | located_in P granule | 2 | IDA,IEA | Caenorhabditis_elegans(6239) | Component |
| GO:0043229 | located_in intracellular organelle | 1 | IDA | Caenorhabditis_elegans(6239) | Component |
| GO:0048477 | involved_in oogenesis | 1 | IEA | Caenorhabditis_elegans(6239) | Process |
| GO:1990124 | part_of messenger ribonucleoprotein complex | 1 | IDA | Caenorhabditis_elegans(6239) | Component |
| GO:1990904 | part_of ribonucleoprotein complex | 1 | IDA | Caenorhabditis_elegans(6239) | Component |