| RBP Type |
N.A. |
| Diseases |
|
| Drug |
|
| Main interacting RNAs |
|
| Moonlighting functions |
|
| Localizations |
|
|
BulkPerturb-seq
|
|
| Ensembl ID | ENSPSMG00000003252 | Gene ID |
|
Accession |
- |
| Symbol |
ETFBKMT |
Alias |
- |
Full Name |
electron transfer flavoprotein subunit beta lysine methyltransferase |
| Status |
Confidence |
Length |
3222 bases |
Strand |
Minus strand |
| Position |
MPIZ01000962.1 : 627825 - 631046 |
RNA binding domain |
Met_10 , MTS |
| Summary |
N.A. |
RNA binding domains (RBDs)

| Protein |
Domain |
Pfam ID |
E-value |
Domain number |
| ENSPSMP00000004058 |
Met_10 |
PF02475 |
0.00035 |
1 |
| ENSPSMP00000004058 |
MTS |
PF05175 |
6.3e-05 |
2 |
RNA binding proteomes (RBPomes)
| Pubmed ID |
Full Name |
Cell |
Author |
Time |
Doi |
Literatures on RNA binding capacity
| Pubmed ID |
Title |
Author |
Time |
Journal |
| Name |
Transcript ID |
bp |
Protein |
Translation ID |
| ETFBKMT-201 |
ENSPSMT00000004915 |
780 |
259aa |
ENSPSMP00000004058 |
| Pathway ID |
Pathway Name |
Source |
| | |
| ensgID |
Trait |
pValue |
Pubmed ID |
| ensgID | SNP | Chromosome | Position |
Trait | PubmedID | Or or BEAT |
EFO ID |
|---|
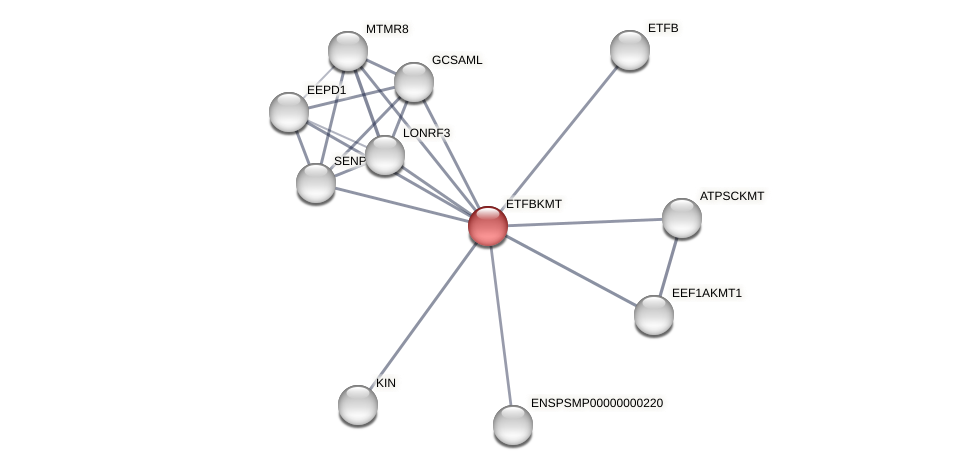
Protein-Protein Interaction (PPI)
| Ensembl ID |
Source |
Target |
| Species |
Protein ID |
Perc_pos |
Perc_id |
Species |
Protein ID |
Perc_pos |
Perc_id |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000007025 | 33.6449 | 17.2897 | prolemur_simus | ENSPSMP00000004058 | 27.7992 | 14.2857 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000021381 | 24.4838 | 12.6844 | prolemur_simus | ENSPSMP00000004058 | 32.0463 | 16.6023 |
| Ensembl ID |
Source |
Target |
| Species |
Protein ID |
Perc_pos |
Perc_id |
Species |
Protein ID |
Perc_pos |
Perc_id |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 45.1737 | 39.3822 | notamacropus_eugenii | ENSMEUP00000003928 | 81.25 | 70.8333 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 59.0734 | 55.5985 | dasypus_novemcinctus | ENSDNOP00000001343 | 93.2927 | 87.8049 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 87.6448 | callithrix_jacchus | ENSCJAP00000065141 | 91.9847 | 86.6412 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.8224 | 89.9614 | cercocebus_atys | ENSCATP00000040967 | 92.7481 | 88.9313 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.8224 | 89.1892 | macaca_fascicularis | ENSMFAP00000042410 | 92.7481 | 88.1679 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.8224 | 89.9614 | macaca_mulatta | ENSMMUP00000014636 | 92.7481 | 88.9313 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.8224 | 89.5753 | macaca_nemestrina | ENSMNEP00000001319 | 92.7481 | 88.5496 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 70.6564 | 63.7066 | papio_anubis | ENSPANP00000058369 | 69.5817 | 62.7376 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 89.5753 | mandrillus_leucophaeus | ENSMLEP00000040280 | 92.3664 | 88.5496 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 88.8031 | gorilla_gorilla | ENSGGOP00000003656 | 91.9847 | 87.7863 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 89.1892 | pan_paniscus | ENSPPAP00000009353 | 92.3664 | 88.1679 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 89.1892 | pan_troglodytes | ENSPTRP00000069758 | 92.3664 | 88.1679 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 88.8031 | pongo_abelii | ENSPPYP00000033557 | 91.9847 | 87.7863 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 88.417 | homo_sapiens | ENSP00000350353 | 91.6031 | 87.4046 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 86.4865 | canis_lupus_familiaris | ENSCAFP00845025652 | 90.2256 | 84.2105 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 86.8726 | vulpes_vulpes | ENSVVUP00000016983 | 90.2256 | 84.5865 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 85.3282 | ursus_americanus | ENSUAMP00000006263 | 90.6015 | 83.0827 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 86.1004 | ailuropoda_melanoleuca | ENSAMEP00000009503 | 90.6015 | 83.8346 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 86.1004 | mustela_putorius_furo | ENSMPUP00000000208 | 90.6015 | 83.8346 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.1197 | 84.1699 | felis_catus | ENSFCAP00000032718 | 88.7218 | 81.9549 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.1197 | 82.6255 | panthera_leo | ENSPLOP00000025463 | 88.7218 | 80.4511 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.1197 | 83.7838 | panthera_pardus | ENSPPRP00000013540 | 88.7218 | 81.5789 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.5058 | 85.7143 | tursiops_truncatus | ENSTTRP00000004588 | 90.458 | 84.7328 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.278 | 86.8726 | delphinapterus_leucas | ENSDLEP00000019994 | 89.8496 | 84.5865 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 86.8726 | physeter_catodon | ENSPCTP00005015889 | 90.2256 | 84.5865 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.8919 | 87.6448 | balaenoptera_musculus | ENSBMSP00010017653 | 90.8397 | 86.6412 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.8919 | 85.7143 | loxodonta_africana | ENSLAFP00000020395 | 89.4737 | 83.4586 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 87.2587 | equus_caballus | ENSECAP00000054974 | 80.602 | 75.5853 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 47.8764 | 45.9459 | equus_asinus | ENSEASP00005049188 | 96.875 | 92.9688 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 89.1892 | 82.2394 | bos_taurus | ENSBTAP00000001167 | 86.8421 | 80.0752 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 90.3475 | 86.1004 | camelus_dromedarius | ENSCDRP00005020420 | 87.9699 | 83.8346 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.1197 | 84.1699 | marmota_marmota_marmota | ENSMMMP00000025109 | 90.0763 | 83.2061 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 90.7336 | 84.1699 | urocitellus_parryii | ENSUPAP00010017581 | 89.6947 | 83.2061 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 87.2587 | 79.5367 | cricetulus_griseus_chok1gshd | ENSCGRP00001018410 | 88.6274 | 80.7843 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 88.8031 | 80.3089 | mesocricetus_auratus | ENSMAUP00000004203 | 90.1961 | 81.5686 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 86.8726 | 78.3784 | mus_caroli | MGP_CAROLIEiJ_P0076973 | 88.2353 | 79.6078 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 86.8726 | 78.3784 | mus_spretus | MGP_SPRETEiJ_P0080246 | 88.2353 | 79.6078 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 86.8726 | 78.3784 | mus_musculus | ENSMUSP00000136167 | 88.2353 | 79.6078 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 86.8726 | 78.3784 | mus_pahari | MGP_PahariEiJ_P0053317 | 88.2353 | 79.6078 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 85.7143 | 78.3784 | rattus_norvegicus | ENSRNOP00000052309 | 87.0588 | 79.6078 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 76.834 | 69.8842 | cavia_porcellus | ENSCPOP00000003567 | 78.3465 | 71.2598 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.0502 | 85.3282 | ursus_maritimus | ENSUMAP00000002994 | 90.6015 | 83.0827 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 89.1892 | 81.4672 | bos_grunniens | ENSBGRP00000007217 | 86.8421 | 79.3233 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 88.8031 | 81.8533 | bos_indicus_hybrid | ENSBIXP00005025536 | 86.4662 | 79.6992 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 94.5946 | 90.3475 | otolemur_garnettii | ENSOGAP00000006772 | 94.2308 | 90 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 56.7568 | 54.0541 | vicugna_pacos | ENSVPAP00000004058 | 93.6306 | 89.172 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 57.9151 | 54.4402 | tupaia_belangeri | ENSTBEP00000010539 | 95.5414 | 89.8089 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 87.6448 | aotus_nancymaae | ENSANAP00000021646 | 92.3664 | 86.6412 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 58.6873 | 54.4402 | saimiri_boliviensis_boliviensis | ENSSBOP00000009859 | 96.8153 | 89.8089 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 87.6448 | monodon_monoceros | ENSMMNP00015026242 | 90.2256 | 85.3383 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.1197 | 84.556 | ictidomys_tridecemlineatus | ENSSTOP00000020105 | 90.0763 | 83.5878 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 89.1892 | 82.2394 | bison_bison_bison | ENSBBBP00000011932 | 86.8421 | 80.0752 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.278 | 85.3282 | sciurus_vulgaris | ENSSVLP00005024898 | 91.2214 | 84.3511 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 88.8031 | rhinolophus_ferrumequinum | ENSRFEP00010005605 | 90.9774 | 86.4662 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.8224 | 89.5753 | chlorocebus_sabaeus | ENSCSAP00000003679 | 92.7481 | 88.5496 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 89.1892 | rhinopithecus_bieti | ENSRBIP00000017431 | 92.3664 | 88.1679 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 89.5753 | 83.7838 | moschus_moschiferus | ENSMMSP00000020791 | 75.57 | 70.684 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 89.1892 | rhinopithecus_roxellana | ENSRROP00000021162 | 92.3664 | 88.1679 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 89.1892 | 81.4672 | bos_mutus | ENSBMUP00000017216 | 86.8421 | 79.3233 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.278 | 88.417 | nomascus_leucogenys | ENSNLEP00000020040 | 91.5709 | 87.7395 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 88.417 | 79.9228 | nannospalax_galili | ENSNGAP00000024909 | 89.8039 | 81.1765 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.8919 | 85.7143 | pteropus_vampyrus | ENSPVAP00000013429 | 89.4737 | 83.4586 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 96.139 | 94.2085 | propithecus_coquereli | ENSPCOP00000002867 | 95.4023 | 93.4866 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 86.4865 | canis_lupus_dingo | ENSCAFP00020024249 | 90.2256 | 84.2105 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 92.6641 | 85.7143 | neovison_vison | ENSNVIP00000026082 | 90.2256 | 83.4586 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 91.5058 | 84.1699 | panthera_tigris_altaica | ENSPTIP00000011351 | 89.0977 | 81.9549 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 93.4363 | 87.6448 | cebus_imitator | ENSCCAP00000029899 | 92.3664 | 86.6412 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 15.0579 | 7.33591 | ciona_intestinalis | ENSCINP00000014533 | 43.3333 | 21.1111 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 42.8571 | 31.2741 | cyprinodon_variegatus | ENSCVAP00000028543 | 62.3596 | 45.5056 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 52.8958 | 38.2239 | neolamprologus_brichardi | ENSNBRP00000019000 | 59.3074 | 42.8571 |
| ENSPSMG00000003252 | prolemur_simus | ENSPSMP00000004058 | 40.5405 | 31.2741 | labrus_bergylta | ENSLBEP00000019881 | 66.0377 | 50.9434 |
| Go ID |
Go term |
No. evidence |
Entries |
Species |
Category |